Current Biotechnology ›› 2024, Vol. 14 ›› Issue (6): 1042-1054.DOI: 10.19586/j.2095-2341.2024.0092
• Articles • Previous Articles Next Articles
Variation Analysis of SARS-CoV-2 Omicron RdRp and Construction of its Active Reaction System
Xi KANG( ), Ziyu LIN, Yishu YANG, Jintao LI, Minglian WANG(
), Ziyu LIN, Yishu YANG, Jintao LI, Minglian WANG( )
)
- College of Chemistry and Life Sciences,Beijing University of Technology,Beijing 100124,China
-
Received:2024-04-29Accepted:2024-05-22Online:2024-11-25Published:2024-12-27 -
Contact:Minglian WANG
新型冠状病毒Omicron RdRp的变异分析及其活性反应体系构建
- 北京工业大学化学与生命科学学院,北京 100124
-
通讯作者:王明连 -
作者简介:康茜 E-mail: kangx@emails.bjut.edu.cn; -
基金资助:北京市自然科学基金资助项目(M22032)
CLC Number:
Cite this article
Xi KANG, Ziyu LIN, Yishu YANG, Jintao LI, Minglian WANG. Variation Analysis of SARS-CoV-2 Omicron RdRp and Construction of its Active Reaction System[J]. Current Biotechnology, 2024, 14(6): 1042-1054.
康茜, 林姿妤, 杨怡姝, 李劲涛, 王明连. 新型冠状病毒Omicron RdRp的变异分析及其活性反应体系构建[J]. 生物技术进展, 2024, 14(6): 1042-1054.
share this article
| 分支 | 低频错义突变位点 | 序列号 |
|---|---|---|
| BA.2 | N507I、F694Y | EPI_ISL_16923349 |
| BA.2.12.1 | A253V | EPI_ISL_12812554 |
| S433G | EPI_ISL_14226909 | |
| I696V | EPI_ISL_12315615 | |
| BA.5 | N88S | C_AA011039.1、C_AA021207.1、EPI_ISL_17775683 |
| T141I | EPI_ISL_15679848 | |
| L247F | C_AA003174.1、C_AA003375.1、C_AA007149.1、C_AA007379.1、C_AA007675.1、C_AA007687.1、C_AA008584.1、C_AA012664.1、C_AA013792.1、C_AA015837.1、EPI_ISL_16634016、EPI_ISL_16709169、EPI_ISL_16955401、EPI_ISL_17199703、EPI_ISL_17307371、EPI_ISL_17490482、EPI_ISL_17775462 | |
| L247F、T739I | C_AA010715.1 | |
| V257F | EPI_ISL_16327678 | |
| M463V | EPI_ISL_13140517 | |
| V473F | C_AA001486.1 | |
| Y479H、F480L | C_AA005712.1 | |
| Y521C | EPI_ISL_16327492 | |
| R651C | EPI_ISL_16854471 | |
| G672S | C_AA037046.1 | |
| A771V | EPI_ISL_17581363 | |
| Q822H | EPI_ISL_16922574 | |
| XBB.1.5 | G671S、D63N | EPI_ISL_17999279 |
| G671S、K160N | C_AA035703.1 | |
| G671S、W162C | EPI_ISL_18060791 | |
| G671S、P169S | EPI_ISL_17770420、EPI_ISL_18060868 | |
| G671S、A250V | EPI_ISL_18208365 | |
| G671S、A250V、T806I | C_AA041899.1 | |
| G671S、V299F | EPI_ISL_18040229 | |
| G671S、F317L | EPI_ISL_17762877 | |
| G671S、V320M | EPI_ISL_17816964 | |
| G671S、N734S | EPI_ISL_17696976、EPI_ISL_18105830、EPI_ISL_18161005、EPI_ISL_18217155 | |
| G671S、N743D | C_AA017934.1、C_AA026969.1、C_AA027243.1、C_AA027499.1、C_AA031571.1、EPI_ISL_17735939、EPI_ISL_17801950、EPI_ISL_17808433、EPI_ISL_17836704、EPI_ISL_17978440 | |
| XBB.1.16 | G671S、S6P | C_AA032258.1 |
| G671S、D40A | C_AA037478.1、EPI_ISL_18074400 | |
| G671S、A46S | C_AA030394.1 | |
| G671S、K91R | C_AA026796.1、EPI_ISL_18074225 | |
| G671S、N198T | C_AA037503.1 | |
| G671S、T226M | C_AA041901.1 | |
| G671S、T226M、V848I | EPI_ISL_18208356 | |
| G671S、T248I | C_AA029090.1、C_AA038338.1 | |
| G671S、T252I | EPI_ISL_18208460 | |
| G671S、T262K | C_AA036945.1 | |
| G671S、T293I | C_AA014710.1、C_AA027341.1、C_AA033549.1、C_AA042222.1、C_AA043434.1、C_AA043493.1、EPI_ISL_18208043、EPI_ISL_18213311 | |
| G671S、T293I、V848I | C_AA042222.1 | |
| G671S、N781S | C_AA038411.1、EPI_ISL_18105657 | |
| G671S、G823S | C_AA030861.1 | |
| G671S、V848I | C_AA014495.1、EPI_ISL_17736476 | |
| G671S、H882R | C_AA013559.1 | |
| G671S、R914K | C_AA043432.1 | |
| XBB.1.9 | G671S、T586A | C_AA018119.1 |
| EG.5.1 | G671S、D63N、A97V | EPI_ISL_18208463 |
| G671S、D63N、V128I | EPI_ISL_18166323 | |
| G671S、D63N、L323F | EPI_ISL_18208580 | |
| G671S、D63N、V848I | C_AA027550.1 | |
| G671S、N64D | EPI_ISL_18066441 |
Table 1 All low frequency missense mutations in the amino acid sequences of RdRp from Omicron
| 分支 | 低频错义突变位点 | 序列号 |
|---|---|---|
| BA.2 | N507I、F694Y | EPI_ISL_16923349 |
| BA.2.12.1 | A253V | EPI_ISL_12812554 |
| S433G | EPI_ISL_14226909 | |
| I696V | EPI_ISL_12315615 | |
| BA.5 | N88S | C_AA011039.1、C_AA021207.1、EPI_ISL_17775683 |
| T141I | EPI_ISL_15679848 | |
| L247F | C_AA003174.1、C_AA003375.1、C_AA007149.1、C_AA007379.1、C_AA007675.1、C_AA007687.1、C_AA008584.1、C_AA012664.1、C_AA013792.1、C_AA015837.1、EPI_ISL_16634016、EPI_ISL_16709169、EPI_ISL_16955401、EPI_ISL_17199703、EPI_ISL_17307371、EPI_ISL_17490482、EPI_ISL_17775462 | |
| L247F、T739I | C_AA010715.1 | |
| V257F | EPI_ISL_16327678 | |
| M463V | EPI_ISL_13140517 | |
| V473F | C_AA001486.1 | |
| Y479H、F480L | C_AA005712.1 | |
| Y521C | EPI_ISL_16327492 | |
| R651C | EPI_ISL_16854471 | |
| G672S | C_AA037046.1 | |
| A771V | EPI_ISL_17581363 | |
| Q822H | EPI_ISL_16922574 | |
| XBB.1.5 | G671S、D63N | EPI_ISL_17999279 |
| G671S、K160N | C_AA035703.1 | |
| G671S、W162C | EPI_ISL_18060791 | |
| G671S、P169S | EPI_ISL_17770420、EPI_ISL_18060868 | |
| G671S、A250V | EPI_ISL_18208365 | |
| G671S、A250V、T806I | C_AA041899.1 | |
| G671S、V299F | EPI_ISL_18040229 | |
| G671S、F317L | EPI_ISL_17762877 | |
| G671S、V320M | EPI_ISL_17816964 | |
| G671S、N734S | EPI_ISL_17696976、EPI_ISL_18105830、EPI_ISL_18161005、EPI_ISL_18217155 | |
| G671S、N743D | C_AA017934.1、C_AA026969.1、C_AA027243.1、C_AA027499.1、C_AA031571.1、EPI_ISL_17735939、EPI_ISL_17801950、EPI_ISL_17808433、EPI_ISL_17836704、EPI_ISL_17978440 | |
| XBB.1.16 | G671S、S6P | C_AA032258.1 |
| G671S、D40A | C_AA037478.1、EPI_ISL_18074400 | |
| G671S、A46S | C_AA030394.1 | |
| G671S、K91R | C_AA026796.1、EPI_ISL_18074225 | |
| G671S、N198T | C_AA037503.1 | |
| G671S、T226M | C_AA041901.1 | |
| G671S、T226M、V848I | EPI_ISL_18208356 | |
| G671S、T248I | C_AA029090.1、C_AA038338.1 | |
| G671S、T252I | EPI_ISL_18208460 | |
| G671S、T262K | C_AA036945.1 | |
| G671S、T293I | C_AA014710.1、C_AA027341.1、C_AA033549.1、C_AA042222.1、C_AA043434.1、C_AA043493.1、EPI_ISL_18208043、EPI_ISL_18213311 | |
| G671S、T293I、V848I | C_AA042222.1 | |
| G671S、N781S | C_AA038411.1、EPI_ISL_18105657 | |
| G671S、G823S | C_AA030861.1 | |
| G671S、V848I | C_AA014495.1、EPI_ISL_17736476 | |
| G671S、H882R | C_AA013559.1 | |
| G671S、R914K | C_AA043432.1 | |
| XBB.1.9 | G671S、T586A | C_AA018119.1 |
| EG.5.1 | G671S、D63N、A97V | EPI_ISL_18208463 |
| G671S、D63N、V128I | EPI_ISL_18166323 | |
| G671S、D63N、L323F | EPI_ISL_18208580 | |
| G671S、D63N、V848I | C_AA027550.1 | |
| G671S、N64D | EPI_ISL_18066441 |
| RdRp模型 | 抑制剂 | 最低结合能/(kcal·mol-1) | 氢键个数 | 涉及氨基酸残基 |
|---|---|---|---|---|
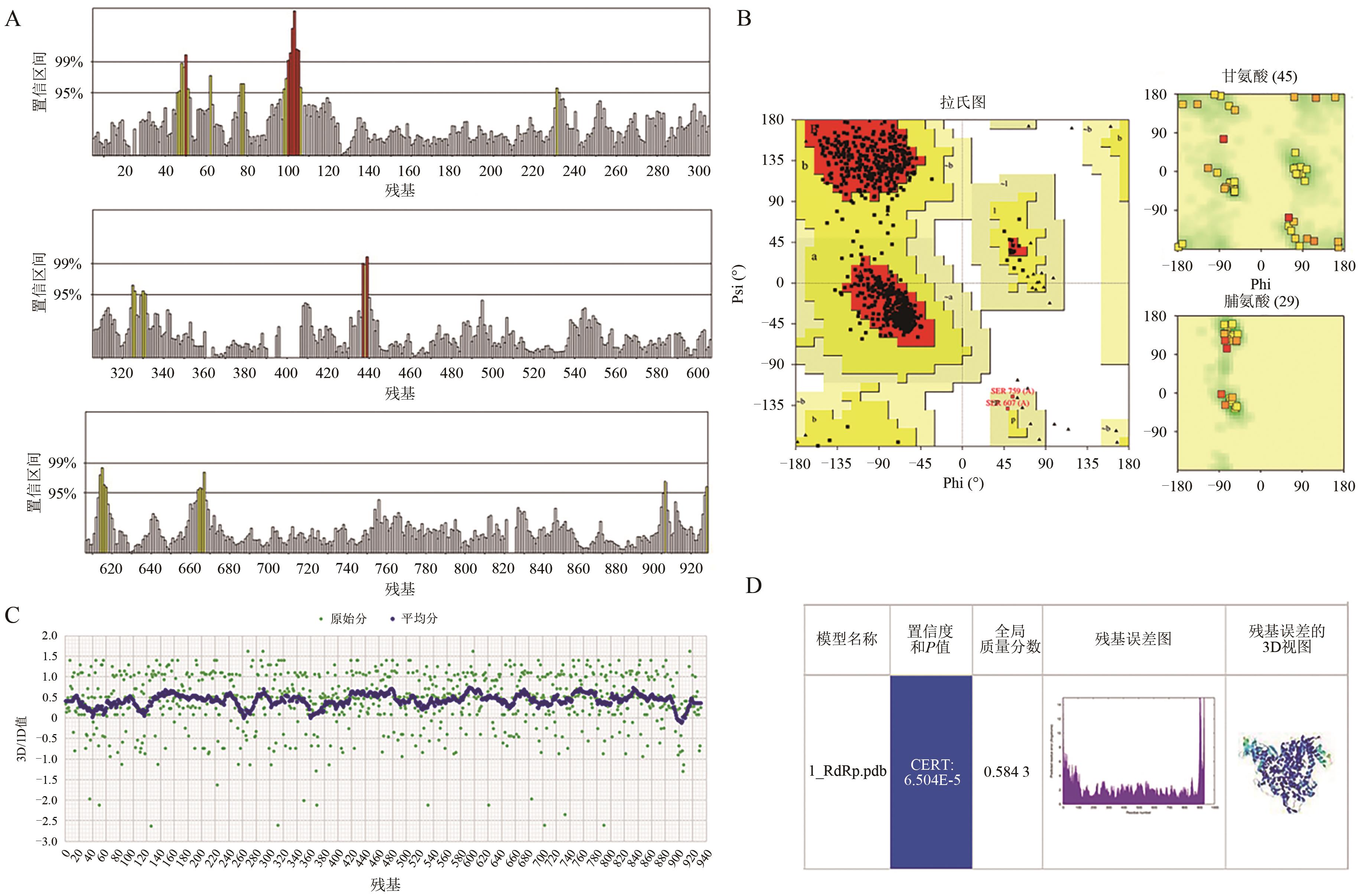
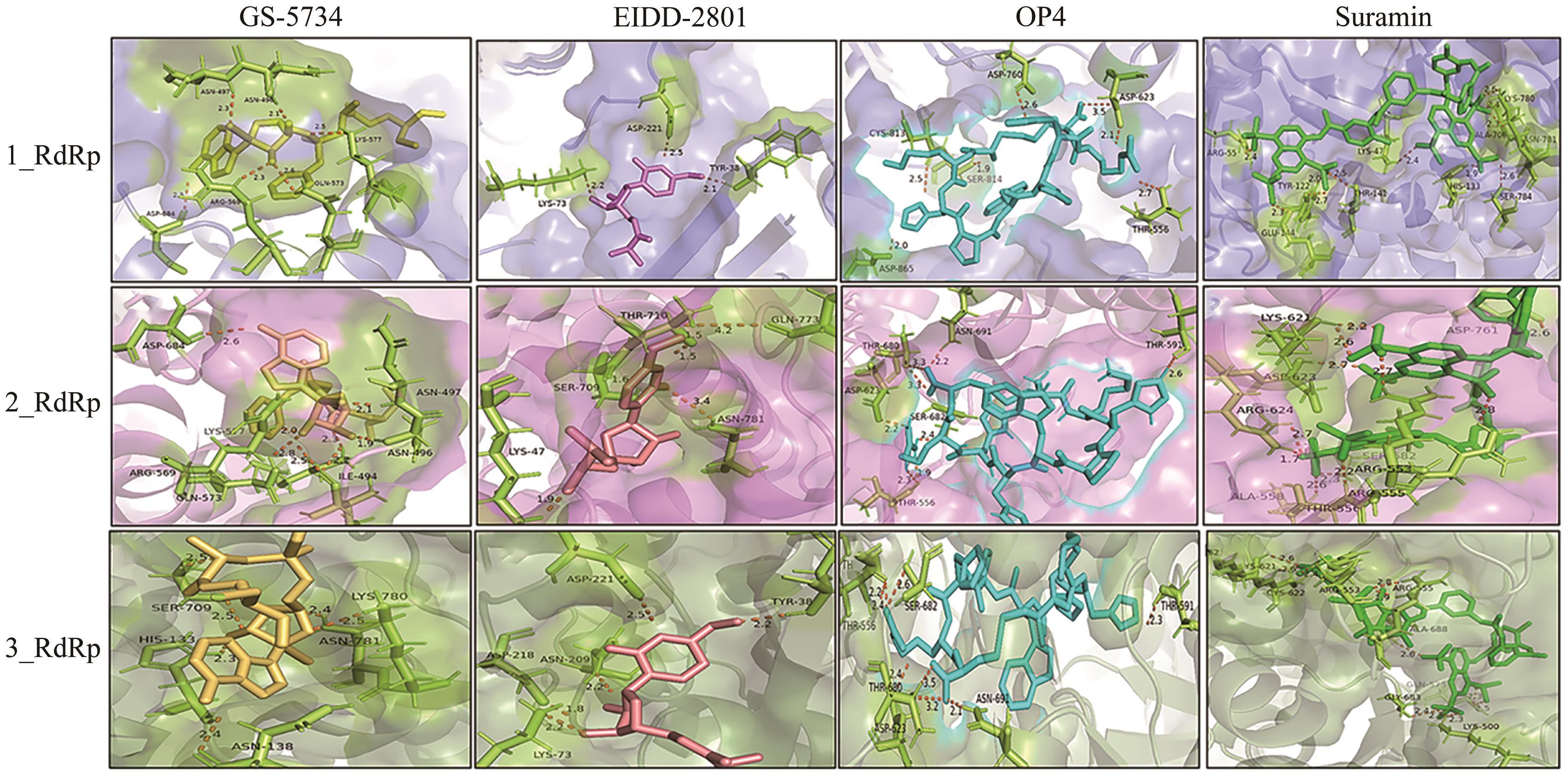
| 1_RdRp | GS-5734 | -7.2 | 6 | Asn496、Asn497、Arg569、Gln573、Lys577、Asp684 |
| OP4 | -8.2 | 7 | Thr556、Asp623、Asp760、Cys813、Ser814、Asp865 | |
| EIDD-2801 | -7.0 | 3 | Tyr38、Lys73、Asp221 | |
| Suramin | -11.1 | 14 | Tyr456、Asn496、Arg555、Thr556、Arg569、Gln573、Lys577、Arg624、Thr680、Ser682、Ala685 | |
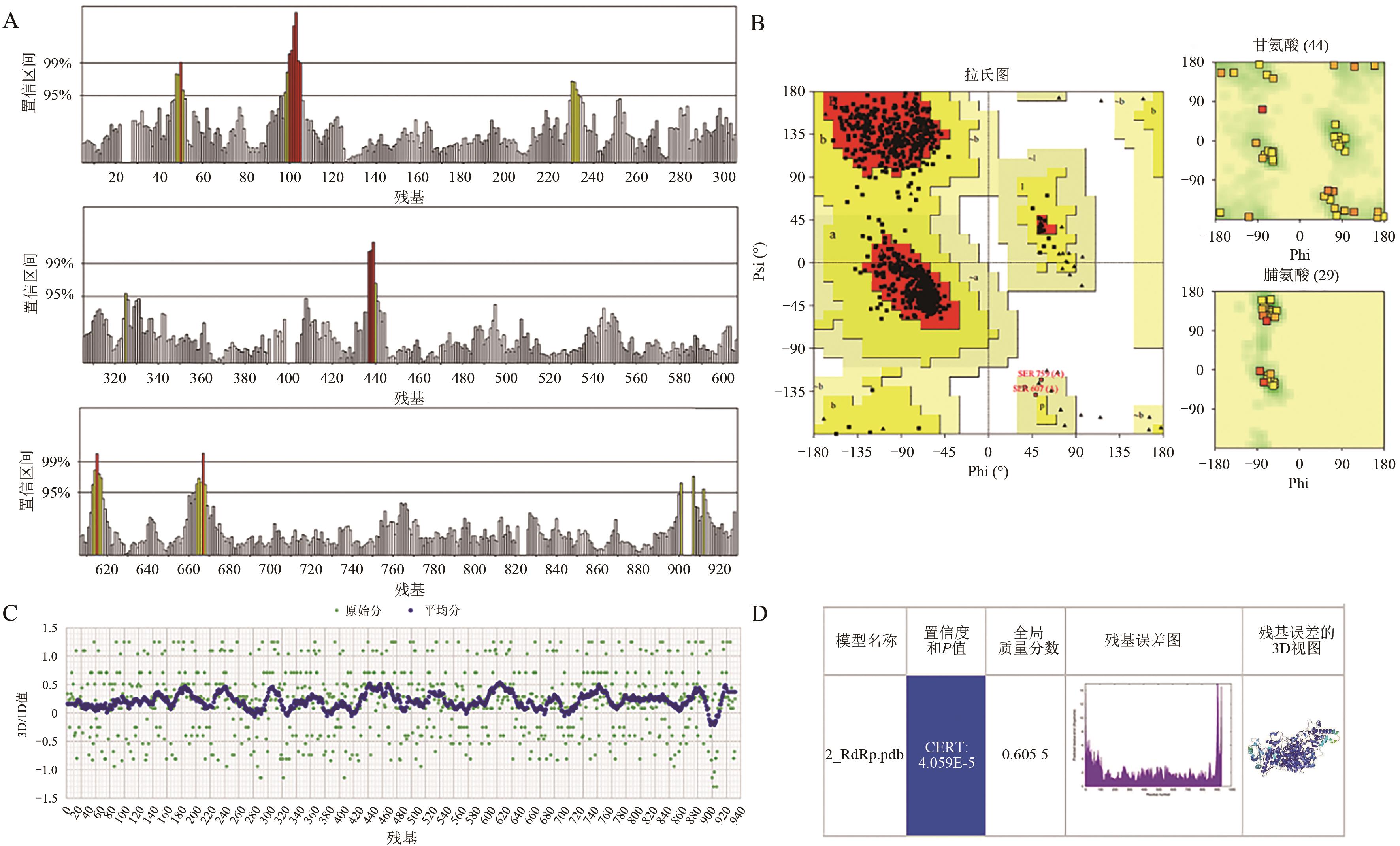
| 2_RdRp | GS-5734 | -7.1 | 8 | Ile494、Asn496、Asn497、Arg569、Gln573、Lys577、Asp684 |
| OP4 | -8.1 | 8 | Thr556、Thr591、Asp623、Thr680、Ser682、Asn691 | |
| EIDD-2801 | -7.2 | 6 | Lys47、Ser709、Thr710、Gln773、Asn781 | |
| Suramin | -11.1 | 11 | Arg553、Arg555、Thr556、Ala558、Lys621、Asp623、Arg624、Ser682、Asp761 | |
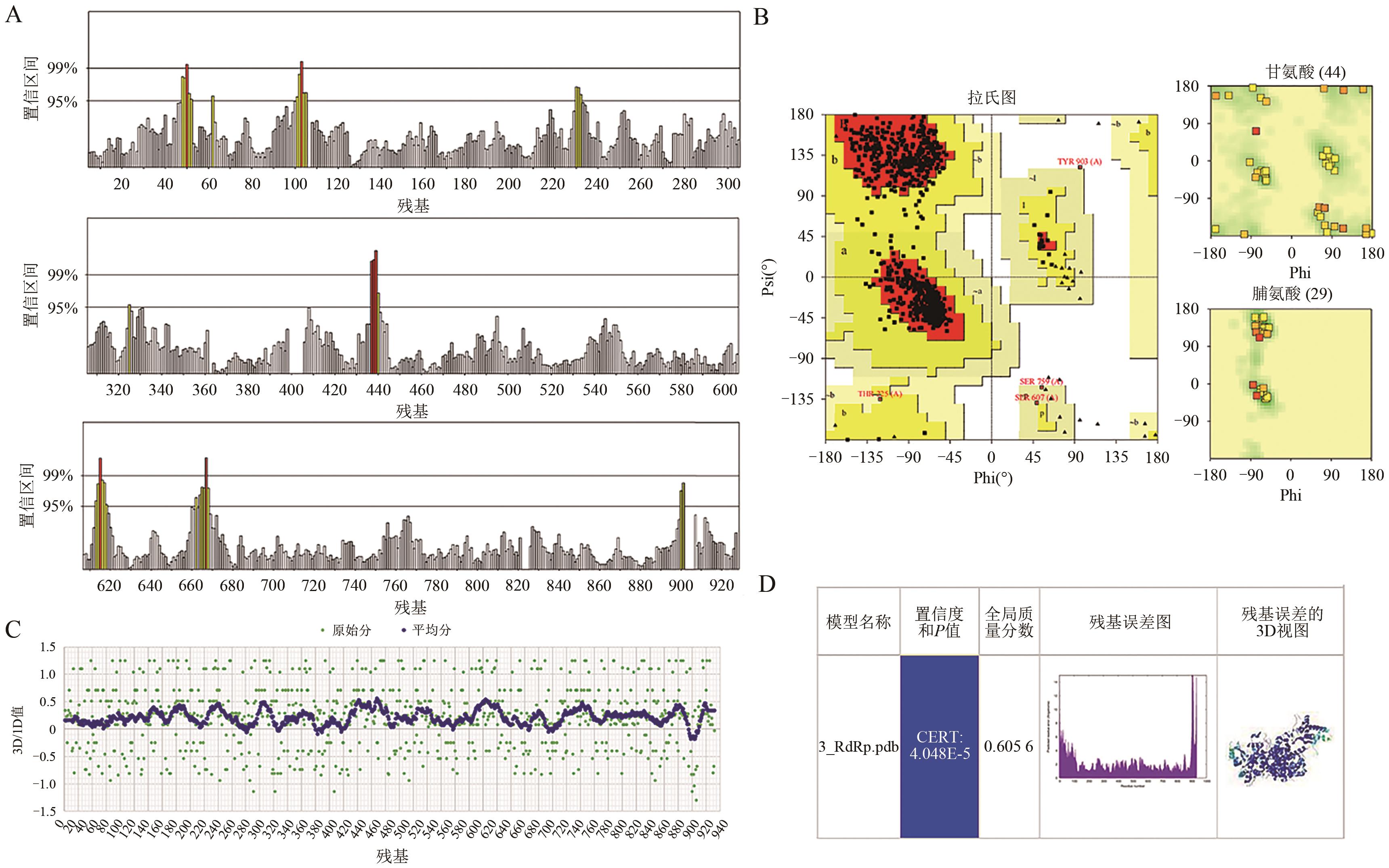
| 3_RdRp | GS-5734 | -7.6 | 6 | His133、Asn138、Ser709、Asn781、Lys780 |
| OP4 | -8.0 | 8 | Thr556、Thr591、Asp623、Thr680、Ser682、Asn691 | |
| EIDD-2801 | -7.1 | 5 | Tyr38、Lys73、Asn209、Asp218、Asp221 | |
| Suramin | -10.8 | 11 | Lys500、Arg553、Arg555、Gln573、Lys621、Cys622、Gly683、Ala688 |
Table 2 Minimum binding energy and interacting amino acid residues of Omicron RdRp docking with four inhibitors
| RdRp模型 | 抑制剂 | 最低结合能/(kcal·mol-1) | 氢键个数 | 涉及氨基酸残基 |
|---|---|---|---|---|
| 1_RdRp | GS-5734 | -7.2 | 6 | Asn496、Asn497、Arg569、Gln573、Lys577、Asp684 |
| OP4 | -8.2 | 7 | Thr556、Asp623、Asp760、Cys813、Ser814、Asp865 | |
| EIDD-2801 | -7.0 | 3 | Tyr38、Lys73、Asp221 | |
| Suramin | -11.1 | 14 | Tyr456、Asn496、Arg555、Thr556、Arg569、Gln573、Lys577、Arg624、Thr680、Ser682、Ala685 | |
| 2_RdRp | GS-5734 | -7.1 | 8 | Ile494、Asn496、Asn497、Arg569、Gln573、Lys577、Asp684 |
| OP4 | -8.1 | 8 | Thr556、Thr591、Asp623、Thr680、Ser682、Asn691 | |
| EIDD-2801 | -7.2 | 6 | Lys47、Ser709、Thr710、Gln773、Asn781 | |
| Suramin | -11.1 | 11 | Arg553、Arg555、Thr556、Ala558、Lys621、Asp623、Arg624、Ser682、Asp761 | |
| 3_RdRp | GS-5734 | -7.6 | 6 | His133、Asn138、Ser709、Asn781、Lys780 |
| OP4 | -8.0 | 8 | Thr556、Thr591、Asp623、Thr680、Ser682、Asn691 | |
| EIDD-2801 | -7.1 | 5 | Tyr38、Lys73、Asn209、Asp218、Asp221 | |
| Suramin | -10.8 | 11 | Lys500、Arg553、Arg555、Gln573、Lys621、Cys622、Gly683、Ala688 |
| 1 | DAVIES N G, ABBOTT S, BARNARD R C, et al.. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England[J/OL]. Science, 2021, 372(6538): eabg3055[2024-03-25]. . |
| 2 | TEGALLY H, WILKINSON E, GIOVANETTI M, et al.. Detection of a SARS-CoV-2 variant of concern in South Africa[J]. Nature, 2021, 592(7854): 438-443. |
| 3 | SABINO E C, BUSS L F, CARVALHO M P S, et al.. Resurgence of COVID-19 in Manaus, Brazil, despite high seroprevalence[J]. Lancet Lond. Engl., 2021, 397(10273): 452-455. |
| 4 | CALLAWAY E. Delta coronavirus variant: scientists brace for impact[J]. Nature, 2021, 595(7865): 17-18. |
| 5 | VIANA R, MOYO S, AMOAKO D G, et al.. Rapid epidemic expansion of the SARS-CoV-2 Omicron variant in southern Africa[J]. Nature, 2022, 603(7902): 679-686. |
| 6 | MEO S A, MEO A S, AL-JASSIR F F, et al.. Omicron SARS-CoV-2 new variant: global prevalence and biological and clinical characteristics[J]. Eur. Rev. Med. Pharmacol. Sci., 2021, 25(24): 8012-8018. |
| 7 | HADFIELD J, MEGILL C, BELL S M, et al.. Nextstrain: real-time tracking of pathogen evolution[J]. Bioinform. Oxf. Engl., 2018, 34(23): 4121-4123. |
| 8 | XIA S, WANG L, ZHU Y, et al.. Origin, virological features, immune evasion and intervention of SARS-CoV-2 Omicron sublineages[J/OL]. Signal Transduct. Target. Ther., 2022, 7(1): 241[2024-03-25]. . |
| 9 | HOSSAIN A, AKTER S, RASHID A A, et al.. Unique mutations in SARS-CoV-2 Omicron subvariants' non-spike proteins: potential impacts on viral pathogenesis and host immune evasion[J/OL]. Microb. Pathog., 2022, 170: 105699[2024-03-25]. . |
| 10 | 孔庆福,马晓洁,刘凤凤,等.2023年6月中国需关注的突发公共卫生事件风险评估[J].疾病监测,2023,38(6):631-635. |
| KONG Q F, MA X J, LIU F F, et al.. Risk assessment of public health emergencies concerned in China, June 2023[J]. Dis. Surveill., 2023, 38(6): 631-635. | |
| 11 | 笃梦雪,严明明,李蔓兮,等.2023年9月中国需关注的突发公共卫生事件风险评估[J].疾病监测,2023,38(9):1025-1028. |
| DU M X, YAN M M, LI M X, et al.. Risk assessment of public health emergencies concerned in China, September 2023[J]. Dis. Surveill., 2023, 38(9): 1025-1028. | |
| 12 | 王盼盼,冯晔囡,白文清,等.2023年11月中国需关注的突发公共卫生事件风险评估[J].疾病监测,2023,38(11):1285-1288. |
| WANG P P, FENG Y N, BAI W Q, et al.. Risk assessment of public health emergencies concerned in China, November 2023[J]. Dis. Surveill., 2023, 38(11): 1285-1288. | |
| 13 | ZHU W, CHEN C Z, GORSHKOV K, et al.. RNA-dependent RNA polymerase as a target for COVID-19 drug discovery[J]. SLAS Discov., 2020, 25(10): 1141-1151. |
| 14 | MAJUMDAR P, NIYOGI S. SARS-CoV-2 mutations: the biological trackway towards viral fitness[J/OL]. Epidemiol. Infect., 2021, 149: e110[2024-03-25]. . |
| 15 | YURKOVETSKIY L, WANG X, PASCAL K E, et al.. Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant[J]. Cell, 2020, 183(3): 739-751. |
| 16 | MALONE B, URAKOVA N, SNIJDER E J, et al.. Structures and functions of coronavirus replication-transcription complexes and their relevance for SARS-CoV-2 drug design[J]. Nat. Rev. Mol. Cell Biol., 2022, 23(1): 21-39. |
| 17 | BAI C, ZHONG Q, GAO G F. Overview of SARS-CoV-2 genome-encoded proteins[J]. Sci. China Life Sci., 2022, 65(2): 280-294. |
| 18 | MIN J S, KIM G-W, KWON S, et al.. A cell-based reporter assay for screening inhibitors of MERS coronavirus RNA-dependent RNA polymerase activity[J/OL]. J. Clin. Med., 2020, 9(8): 2399[2024-03-25]. . |
| 19 | SONG S, MA L, ZOU D, et al.. The global landscape of SARS-CoV-2 genomes, variants, and haplotypes in 2019nCoVR[J]. Genom. Proteom. Bioinform., 2020, 18(6): 749-759. |
| 20 | Members and Partnerscncb-NGDC. Database resources of the national genomics data center, China national center for bioinformation in 2023[J/OL]. Nucleic Acids Res., 2023, 51(D1): D18-D28[2024-03-25]. . |
| 21 | CHEN M, MA Y, WU S, et al.. Genome warehouse: a public repository housing genome-scale data[J]. Genom. Proteom. Bioinform., 2021, 19(4): 584-589. |
| 22 | KHARE S, GURRY C, FREITAS L, et al.. GISAID's role in pandemic response[J]. China CDC Week., 2021, 3(49): 1049-1051. |
| 23 | GAO Y, YAN L, HUANG Y, et al.. Structure of the RNA-dependent RNA polymerase from COVID-19 virus[J]. Science, 2020, 368(6492): 779-782. |
| 24 | COLOVOS C, YEATES T O. Verification of protein structures: patterns of nonbonded atomic interactions[J]. Protein Sci., 1993, 2(9): 1511-1519. |
| 25 | LÜTHY R, BOWIE J U, EISENBERG D. Assessment of protein models with three-dimensional profiles[J]. Nature, 1992, 356(6364): 83-85. |
| 26 | LASKOWSKI R A, RULLMANNN J A, MACARTHUR M W, et al.. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR[J]. J. Biomol. NMR, 1996, 8(4): 477-486. |
| 27 | MCGUFFIN L J, ALDOWSARI F M F, ALHARBI S M A, et al.. ModFOLD8: accurate global and local quality estimates for 3D protein models[J/OL]. Nucleic Acids Res., 2021, 49(W1): W425-W430[2024-03-25]. . |
| 28 | YIN W, MAO C, LUAN X, et al.. Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir[J]. Science, 2020, 368(6498): 1499-1504. |
| 29 | JIANG Y, YIN W, XU H E. RNA-dependent RNA polymerase: Structure, mechanism, and drug discovery for COVID-19[J]. Biochem. Biophys. Res. Commun., 2021, 538: 47-53. |
| 30 | BEIGEL J H, TOMASHEK K M, DODD L E, et al.. Remdesivir for the treatment of covid-19 - final report[J]. N. Engl. J. Med., 2020, 383(19): 1813-1826. |
| 31 | 苏小进,王明连,袁润余,等.登革病毒RdRp配体寡肽潜在广谱靶向作用研究[J].国际病毒学杂志,2022,29(4):286-291. |
| SU X J, WANG M L, YUAN R Y, et al.. Potential broad-spectrum targeting effects of the ligand oligopeptide of dengue virus RdRp[J]. Int. J. Virol., 2022, 29(4): 286-291. | |
| 32 | AGOSTINI M L, PRUIJSSERS A J, CHAPPELL J D, et al.. Small-molecule antiviral β-d-N 4-hydroxycytidine inhibits a proofreading-intact coronavirus with a high genetic barrier to resistance[J]. J. Virol., 2019, 93(24): e01348-19. |
| 33 | FISCHER W A, ERON J J, HOLMAN W, et al.. A phase 2a clinical trial of molnupiravir in patients with COVID-19 shows accelerated SARS-CoV-2 RNA clearance and elimination of infectious virus[J/OL]. Sci. Transl. Med., 2022, 14(628): eabl7430 [2024-03-25]. . |
| 34 | YIN W, LUAN X, LI Z, et al.. Structural basis for inhibition of the SARS-CoV-2 RNA polymerase by suramin[J]. Nat. Struct. Mol. Biol., 2021, 28(3): 319-325. |
| 35 | WU F, ZHAO S, YU B, et al.. A new coronavirus associated with human respiratory disease in China[J]. Nature, 2020, 579(7798): 265-269. |
| 36 | 杨高松,马东杰.呼吸道传染病治疗中抗体药物的研发进展[J].生物技术进展,2020,10(5):441-447. |
| YANG G S, MA D J. Progress in research and development of antibodies for treatment of respiratory infectious diseases[J]. Curr. Biotechnol., 2020, 10(5): 441-447. | |
| 37 | 杨懿祺,张志高,游小龙,等.抗体药物的发展与应用[J].生物技术进展,2022,12(3):358-365. |
| YANG Y Q, ZHANG Z G, YOU X L, et al.. Development and application of monoclonal antibody-based drug[J]. Curr. Biotechnol., 2022, 12(3): 358-365. | |
| 38 | MCDONALD S M. RNA synthetic mechanisms employed by diverse families of RNA viruses[J]. Wiley Interdiscip. Rev. RNA, 2013, 4(4): 351-367. |
| 39 | PICARAZZI F, VICENTI I, SALADINI F, et al.. Targeting the RdRp of emerging RNA viruses: the structure-based drug design challenge[J/OL]. Molecules, 2020, 25(23): 5695[2024-03-25]. . |
| 40 | LI Q, YI D, LEI X, et al.. Corilagin inhibits SARS-CoV-2 replication by targeting viral RNA-dependent RNA polymerase[J]. Acta Pharm. Sin. B, 2021, 11(6): 1555-1567. |
| 41 | BAI X, SUN H, WU S, et al.. Identifying small-molecule inhibitors of SARS-CoV-2 RNA-dependent RNA polymerase by establishing a fluorometric assay[J/OL]. Front. Immunol., 2022, 13: 844749[2024-03-25]. . |
| [1] | Yanhua LIU, Rui GUO, Yanyi LI, Jinli CHEN. Progress on the Development of Vaccines Related to Chronic Respiratory Diseases [J]. Current Biotechnology, 2025, 15(3): 396-403. |
| [2] | Yezi MA, Meijuan XIA, Cuicui LIU, Hongtao WANG, Jiaxi ZHOU. Comparative Analysis of Platelet Transcriptome and Proteome Changes in SARS-CoV2 Omicron Infection [J]. Current Biotechnology, 2024, 14(4): 649-656. |
| [3] | Changhu KE, Yaqing WU, Xueru DING, Huimin HUANG, Hui YAN. Research on the Mechanism of Jinyinhua Oral Liquid in the Prevention and Treatment of COVID-19 Based on Network Pharmacology and Molecular Docking [J]. Current Biotechnology, 2023, 13(5): 807-817. |
| [4] | Feng DONG, Bingxin HUANGFU, Jia XU, Wentao XU. Mechanism Exploring of Coix Seed Intervention in Fatty Liver Disease Based on Network Pharmacology [J]. Current Biotechnology, 2023, 13(2): 264-272. |
| [5] | Xiaoyan PENG, Panpan LU, Zuquan HU. Prokaryotic Expression of the RBD Protein of SARS-CoV-2 and its Polyclonal Antibodies Preparation [J]. Current Biotechnology, 2023, 13(1): 102-106. |
| [6] | Haining WANG, Xingjian LIU, Xintao GAO, Yinv LI, Xingjia SHEN, Zhifang ZHANG, Yongzhu YI. Prokaryotic Expression and Neutralization Activity Detection of SARS-CoV-2 Neutralizing Nanobody [J]. Current Biotechnology, 2022, 12(5): 754-759. |
| [7] | HU Sihong, YOU Guoye. Application Prospects of Digital PCR in Detection of SARS-CoV-2 [J]. Curr. Biotech., 2020, 10(6): 674-679. |
| [8] | SUN Jing1, LIU Dan2*, HUI Wenqi2. Study on Design of Colchicine Inhibitors of Tubulin for the Treatment of Rheumatoid Arthritis Based on Computer Simulation [J]. Curr. Biotech., 2019, 9(1): 84-88. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||