


生物技术进展 ›› 2026, Vol. 16 ›› Issue (1): 29-37.DOI: 10.19586/j.2095-2341.2025.0107
收稿日期:2025-08-18
接受日期:2025-10-22
出版日期:2026-01-25
发布日期:2026-02-12
通讯作者:
张正平
作者简介:曾鑫 E-mail: xinzeng@cttq.com;
基金资助:
Xin ZENG( ), Chen WANG, Zhengjun WANG, Hechu LIANG, Zhengping ZHANG(
), Chen WANG, Zhengjun WANG, Hechu LIANG, Zhengping ZHANG( )
)
Received:2025-08-18
Accepted:2025-10-22
Online:2026-01-25
Published:2026-02-12
Contact:
Zhengping ZHANG
摘要:
抗体药物开发面临周期长(>3年)、成本高(>2亿美元)以及多属性协同优化困难等产业瓶颈。传统方法如杂交瘤技术存在通量低、全局优化能力不足等局限。近年来,深度学习(deep learning,DL)技术为抗体药物的智能化开发提供了突破性解决方案。系统综述了DL在抗体药物开发中的研究进展,重点探讨了抗体序列设计、结构预测、亲和力预测与成熟、多目标优化等核心环节的代表性方法与技术挑战,并对未来发展进行了展望,以期为抗体药物研发向智能化、全局化方向转型提供参考。
中图分类号:
曾鑫, 王辰, 王正俊, 梁何楚, 张正平. 抗体药物智能开发的深度学习策略[J]. 生物技术进展, 2026, 16(1): 29-37.
Xin ZENG, Chen WANG, Zhengjun WANG, Hechu LIANG, Zhengping ZHANG. Deep Learning Strategies for Intelligent Development of Antibody Drugs[J]. Current Biotechnology, 2026, 16(1): 29-37.
| 名称 | 类别 | 模型 | 特点 | 参考文献 |
|---|---|---|---|---|
| AbLang | 抗体 | 蛋白质语言模型 | 通过改进的序列-功能关系捕捉能力执行抗体语言建模 | [ |
| AntiBERTy | 抗体 | 蛋白质语言模型 | 基于BERT架构实现抗体序列的全面理解 | [ |
| IgLM | 抗体 | 蛋白质语言模型 | 生成具有更高质量和多样性的抗体序列 | [ |
| nanoBERT | 纳米抗体 | 蛋白质语言模型 | 针对骆驼科VHH结构域设计的专用纳米抗体语言模型 | [ |
| ProGen2-OAS | 抗体 | 蛋白质语言模型 | 针对条件序列生成优化的抗体设计模型 | [ |
| IgBert | 抗体 | 蛋白质语言模型 | 专用于免疫球蛋白序列分析与分类的BERT模型 | [ |
| IgT5 | 抗体 | 蛋白质语言模型 | 基于T5架构的抗体模型,支持翻译、生成和重构任务 | [ |
| GearBind | 抗体 | 图神经网络 | 通过几何注意力机制预测抗体-抗原结合 | [ |
| RefineGNN | 抗体 | 图神经网络 | 使用图神经网络优化抗体序列以提高结合能力 | [ |
| ProteinMPNN | 通用 | 图神经网络 | 基于3D结构条件设计的蛋白质逆向折叠模型(最先进技术) | [ |
| AbDiffuser | 抗体 | 扩散模型 | 基于抗原结构条件生成抗体序列的扩散模型 | [ |
| RFdiffusion | 通用 | 扩散模型 | 具备序列设计能力的蛋白质主链生成模型 | [ |
| Chroma | 通用 | 扩散模型 | 支持结构约束的高分辨率全新蛋白质设计 | [ |
表1 抗体序列设计代表性模型
Table 1 Comparison of representative antibody sequence design models
| 名称 | 类别 | 模型 | 特点 | 参考文献 |
|---|---|---|---|---|
| AbLang | 抗体 | 蛋白质语言模型 | 通过改进的序列-功能关系捕捉能力执行抗体语言建模 | [ |
| AntiBERTy | 抗体 | 蛋白质语言模型 | 基于BERT架构实现抗体序列的全面理解 | [ |
| IgLM | 抗体 | 蛋白质语言模型 | 生成具有更高质量和多样性的抗体序列 | [ |
| nanoBERT | 纳米抗体 | 蛋白质语言模型 | 针对骆驼科VHH结构域设计的专用纳米抗体语言模型 | [ |
| ProGen2-OAS | 抗体 | 蛋白质语言模型 | 针对条件序列生成优化的抗体设计模型 | [ |
| IgBert | 抗体 | 蛋白质语言模型 | 专用于免疫球蛋白序列分析与分类的BERT模型 | [ |
| IgT5 | 抗体 | 蛋白质语言模型 | 基于T5架构的抗体模型,支持翻译、生成和重构任务 | [ |
| GearBind | 抗体 | 图神经网络 | 通过几何注意力机制预测抗体-抗原结合 | [ |
| RefineGNN | 抗体 | 图神经网络 | 使用图神经网络优化抗体序列以提高结合能力 | [ |
| ProteinMPNN | 通用 | 图神经网络 | 基于3D结构条件设计的蛋白质逆向折叠模型(最先进技术) | [ |
| AbDiffuser | 抗体 | 扩散模型 | 基于抗原结构条件生成抗体序列的扩散模型 | [ |
| RFdiffusion | 通用 | 扩散模型 | 具备序列设计能力的蛋白质主链生成模型 | [ |
| Chroma | 通用 | 扩散模型 | 支持结构约束的高分辨率全新蛋白质设计 | [ |
| 名称 | 模型 | 优势 | 局限 | 应用 | 参考文献 |
|---|---|---|---|---|---|
| ABodyBuilder2 | AF-Multimer | 减少OpenMM物理约束问题 | CDR H3结构预测及物理合理结构生成仍具挑战 | Fv及纳米抗体结构预测 | [ |
| tFold-Ab | AF-Multimer | 结合蛋白质语言模型与AF-Multimer | PLM选择需进一步研究 | Fv及纳米抗体结构预测 | [ |
| xTrimoABFold | AF-Multimer | 采用AntiBERTy嵌入与快速模板搜索算法 | 未考虑复合物结构优化 | Fv及纳米抗体结构预测 | [ |
| ABlooper | E(n)-EGNNs | 无需外部工具/MSA,速度快 | 可能产生非物理预测 | CDR环结构预测 | [ |
| DeepAb | LSTM+residual NN | 支持点突变建议 | 依赖Rosetta(速度较慢) | Fv结构预测 | [ |
| IgFold | AntiBERTy+Graph transformer+IPA | AntiBERTy嵌入减少物理约束 | 依赖Rosetta生成主干结构 | Fv及纳米抗体结构预测 | [ |
| EquiFold | SE(3)-equivariant NN | 无需MSA/PLM,参数少速度快 | 可能产生非物理预测 | 迷你蛋白设计及Fv结构预测 | [ |
表2 基于深度学习的抗体结构预测模型
Table 2 Deep learning-based models for antibody structure prediction
| 名称 | 模型 | 优势 | 局限 | 应用 | 参考文献 |
|---|---|---|---|---|---|
| ABodyBuilder2 | AF-Multimer | 减少OpenMM物理约束问题 | CDR H3结构预测及物理合理结构生成仍具挑战 | Fv及纳米抗体结构预测 | [ |
| tFold-Ab | AF-Multimer | 结合蛋白质语言模型与AF-Multimer | PLM选择需进一步研究 | Fv及纳米抗体结构预测 | [ |
| xTrimoABFold | AF-Multimer | 采用AntiBERTy嵌入与快速模板搜索算法 | 未考虑复合物结构优化 | Fv及纳米抗体结构预测 | [ |
| ABlooper | E(n)-EGNNs | 无需外部工具/MSA,速度快 | 可能产生非物理预测 | CDR环结构预测 | [ |
| DeepAb | LSTM+residual NN | 支持点突变建议 | 依赖Rosetta(速度较慢) | Fv结构预测 | [ |
| IgFold | AntiBERTy+Graph transformer+IPA | AntiBERTy嵌入减少物理约束 | 依赖Rosetta生成主干结构 | Fv及纳米抗体结构预测 | [ |
| EquiFold | SE(3)-equivariant NN | 无需MSA/PLM,参数少速度快 | 可能产生非物理预测 | 迷你蛋白设计及Fv结构预测 | [ |
| 名称 | 模型 | 优势 | 局限 | 应用 |
|---|---|---|---|---|
| AlphaFold2(AF2) | Evoformer模块+端到端优化 | 框架区预测精度高;整体建模接近实验分辨率 | CDR-H3预测受限于MSA稀疏性;依赖进化信息 | 单体蛋白预测、全蛋白组建模 |
| AlphaFold3(AF3) | PairFormer+扩散模型,多链复合物预测 | 显著提升复合物、抗体-抗原界面与配体/核酸结合位点的预测精度;在界面残基取向和距离判断上优于AF2 | 计算成本较高;部分复杂体系预测仍不稳定 | 高精度复合物建模、抗体-抗原相互作用预测 |
| ESM-2 | 大规模语言模型(15B 参数),无需 MSA | 序列→结构预测快速;支持大规模抗体库建模 | 对环区/复合物预测精度有限 | 快速抗体结构生成、高通量筛选 |
| ESM-3 | 多模态生成模型(序列-结构-功能联合建模) | 可设计新型功能蛋白;支持可控生成 | 模型参数巨大,资源需求高;完整版本未全面开源 | 抗体功能优化、序列-结构-功能一体化设计 |
表3 通用蛋白质折叠模型的比较
Table 3 Comparison of general protein folding models
| 名称 | 模型 | 优势 | 局限 | 应用 |
|---|---|---|---|---|
| AlphaFold2(AF2) | Evoformer模块+端到端优化 | 框架区预测精度高;整体建模接近实验分辨率 | CDR-H3预测受限于MSA稀疏性;依赖进化信息 | 单体蛋白预测、全蛋白组建模 |
| AlphaFold3(AF3) | PairFormer+扩散模型,多链复合物预测 | 显著提升复合物、抗体-抗原界面与配体/核酸结合位点的预测精度;在界面残基取向和距离判断上优于AF2 | 计算成本较高;部分复杂体系预测仍不稳定 | 高精度复合物建模、抗体-抗原相互作用预测 |
| ESM-2 | 大规模语言模型(15B 参数),无需 MSA | 序列→结构预测快速;支持大规模抗体库建模 | 对环区/复合物预测精度有限 | 快速抗体结构生成、高通量筛选 |
| ESM-3 | 多模态生成模型(序列-结构-功能联合建模) | 可设计新型功能蛋白;支持可控生成 | 模型参数巨大,资源需求高;完整版本未全面开源 | 抗体功能优化、序列-结构-功能一体化设计 |
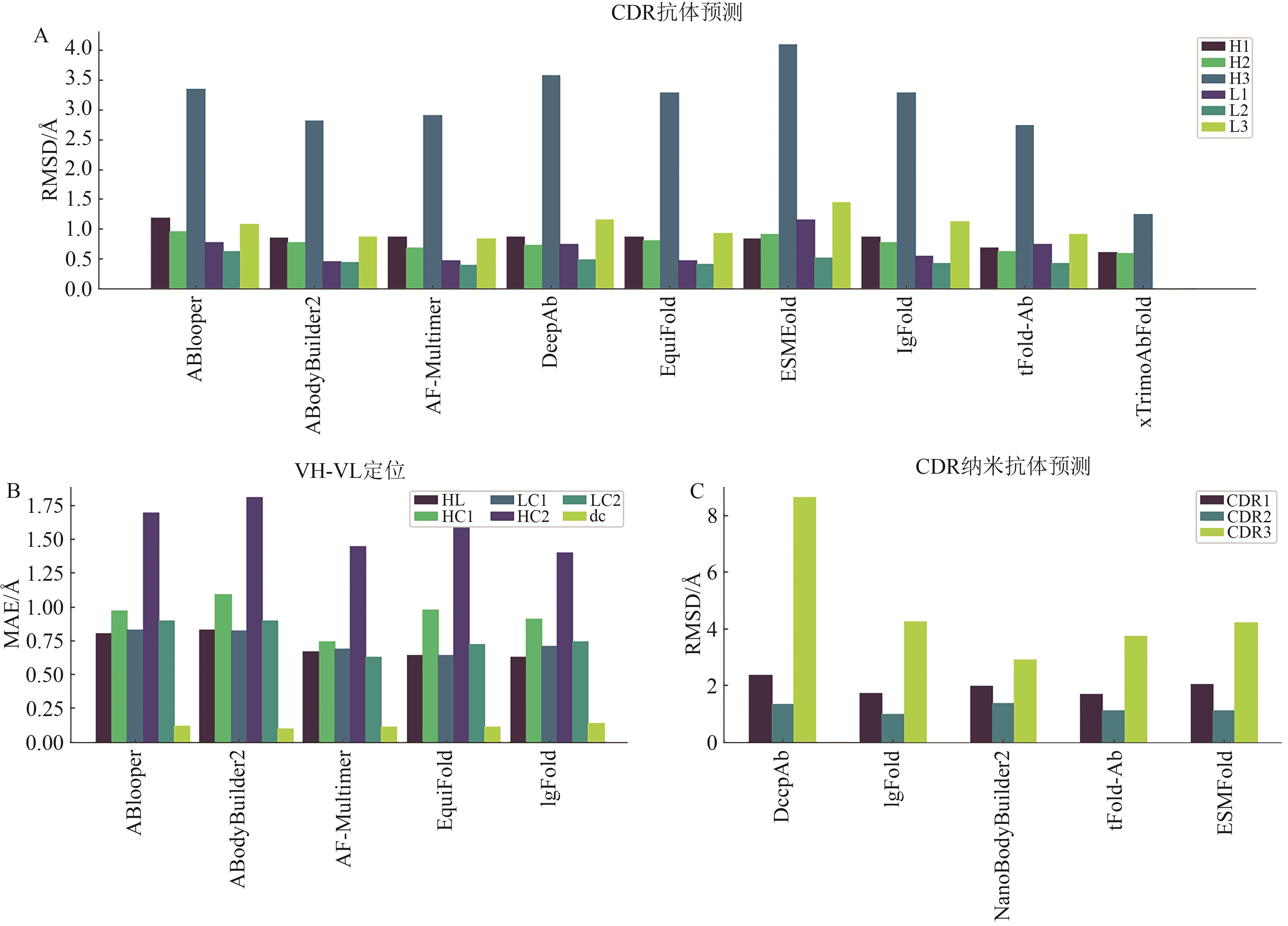
图1 抗体结构预测方法的性能评估[38]A:标准抗体CDR区域预测的RMSD比较;B:VH-VL取向精度MAE比较;C:纳米抗体CDR区域预测的RMSD比较
Fig. 1 Performance evaluation of structural prediction methods for antibodies[38]
| [1] | WANG Z, WANG G, LU H, et al.. Development of therapeutic antibodies for the treatment of diseases[J/OL]. Mol. Biomed., 2022, 3(1): 35[2025-12-21]. . |
| [2] | DEWAKER V, MORYA V K, KIM Y H, et al.. Revolutionizing oncology: the role of artificial intelligence (AI) as an antibody design, and optimization tools[J/OL]. Biomark. Res., 2025, 13(1): 52[2025-12-21]. . |
| [3] | MATSUNAGA R, TSUMOTO K. Accelerating antibody discovery and optimization with high-throughput experimentation and machine learning[J/OL]. J. Biomed. Sci., 2025, 32(1): 46[2025-12-21]. . |
| [4] | 纪思佳,张小雪,翟丽丽,等.抗体药物筛选技术的研究进展[J].生物技术进展,2025,15(1):43-49. |
| JI S J, ZHANG X X, ZHAI L L, et al.. Research progress of antibody drug screening technology[J]. Curr. Biotechnol., 2025, 15(1): 43-49. | |
| [5] | OLSEN T H, MOAL I H, DEANE C M. AbLang: an antibody language model for completing antibody sequences[J/OL]. Bioinform. Adv., 2022, 2: vbac046[2025-12-21]. . |
| [6] | RUFFOLO J A, GRAY J J, SULAM J. Deciphering antibody affinity maturation with language models and weakly supervised learning[EB/OL]. arXiv, 2021: 2112.07782[2025-09-30]. . |
| [7] | SHUAI R W, RUFFOLO J A, GRAY J J. IgLM: infilling language modeling for antibody sequence design[J]. Cell Syst., 2023, 14(11): 979-989. |
| [8] | HADSUND J T, SATŁAWA T, JANUSZ B, et al.. nanoBERT: a deep learning model for gene agnostic navigation of the nanobody mutational space[J/OL]. Bioinform. Adv., 2024, 4(1): vbae033[2025-09-30]. . |
| [9] | NIJKAMP E, RUFFOLO J A, WEINSTEIN E N, et al.. ProGen2: exploring the boundaries of protein language models[J]. Cell Syst., 2023, 14(11): 968-978. |
| [10] | MADANI A, KRAUSE B, GREENE E R, et al.. Large language models generate functional protein sequences across diverse families[J]. Nat. Biotechnol., 2023, 41(8): 1099-1106. |
| [11] | CAI H, ZHANG Z, WANG M, et al.. Pretrainable geometric graph neural network for antibody affinity maturation[J/OL]. Nat. Commun., 2024, 15(1): 7785[2025-12-21]. . |
| [12] | JIN W, WOHLWEND J, BARZILAY R, et al.. Iterative refinement graph neural network for antibody sequence-structure co-design[J/OL]. arXiv, 2021: 2110.04624[2025-09-30]. . |
| [13] | DAUPARAS J, ANISHCHENKO I, BENNETT N, et al.. Robust deep learning-based protein sequence design using ProteinMPNN[J]. Science, 2022, 378(6615): 49-56. |
| [14] | MARTINKUS K, LUDWICZAK J, CHO K, et al.. AbDiffuser: full-atom generation of in vitro functioning antibodies[J/OL]. arXiv, 2023: 2308.05027[2025-09-30]. . |
| [15] | WATSON J L, JUERGENS D, BENNETT N R, et al.. De novo design of protein structure and function with RFdiffusion[J]. Nature, 2023, 620(7976): 1089-1100. |
| [16] | INGRAHAM J B, BARANOV M, COSTELLO Z, et al.. Illuminating protein space with a programmable generative model[J]. Nature, 2023, 623(7989): 1070-1078. |
| [17] | LI Y, LANG Y, XU C, et al.. Benchmarking inverse folding models for antibody CDR sequence design[J/OL]. PLoS One, 2025, 20(6): e0324566[2025-12-21]. . |
| [18] | LIN Z, AKIN H, RAO R, et al.. Evolutionary-scale prediction of atomic-level protein structure with a language model[J]. Science, 2023, 379(6637): 1123-1130. |
| [19] | KENLAY H, DREYER F A, KOVALTSUK A, et al.. Large scale paired antibody language models[J/OL]. PLoS Comput. Biol., 2024, 20(12): e1012646[2025-12-21]. . |
| [20] | WEISSENOW K, ROST B. Are protein language models the new universal key?[J/OL]. Curr. Opin. Struct. Biol., 2025, 91: 102997[2025-12-21]. . |
| [21] | WANG L, LI X, ZHANG H, et al.. A comprehensive review of protein language models[J/OL]. arXiv, 2025: 2502.06881[2025-09-30]. . |
| [22] | BENNETT N R, WATSON J L, RAGOTTE R J, et al.. Atomically accurate de novo design of antibodies with RFdiffusion[J]. Nature, 2025, 649(8095): 183-193. |
| [23] | LUO S, SU Y, PENG X, et al.. Antigen-specific antibody design and optimization with diffusion-based generative models for protein structures[J]. Adv. Neural. Inf. Process Syst., 2022, 35: 9754-9767. |
| [24] | HU Y, TAO F, XU J, et al.. Combining transformer and 3DCNN models to achieve co-design of structures and sequences of antibodies in a diffusional manner[J/OL]. J. Pharm. Anal., 2025, 15(6): 101267[2025-10-21]. . |
| [25] | WU J, KONG X, SUN N, et al.. FlowDesign: improved design of antibody CDRs through flow matching and better prior distributions[J/OL]. Cell Syst., 2025, 16(6): 101270[2025-10-21]. . |
| [26] | ABANADES B, WONG W K, BOYLES F, et al.. ImmuneBuilder: deep-Learning models for predicting the structures of immune proteins[J/OL]. Commun. Biol., 2023, 6(1): 575[2025-12-21]. . |
| [27] | WU J, WU F, JIANG B, et al.. tFold-Ab: fast and accurate antibody structure prediction without sequence homologs[J/OL]. bioRxiv, 2022: 2011-2022[2025-10-21]. . |
| [28] | WANG Y, GONG X, LI S, et al.. xTrimoABFold: de novo antibody structure prediction without MSA[J/OL]. arXiv, 2022: 2212.00735[2025-09-30]. . |
| [29] | ABANADES B, GEORGES G, BUJOTZEK A, et al.. ABlooper: fast accurate antibody CDR loop structure prediction with accuracy estimation[J]. Bioinformatics, 2022, 38(7): 1877-1880. |
| [30] | RUFFOLO J A, SULAM J, GRAY J J. Antibody structure prediction using interpretable deep learning[J/OL]. Patterns, 2022, 3(2): 100406[2025-12-21]. . |
| [31] | RUFFOLO J A, CHU L S, MAHAJAN S P, et al.. Fast, accurate antibody structure prediction from deep learning on massive set of natural antibodies[J/OL]. Nat. Commun., 2023, 14(1): 2389[2025-12-21]. . |
| [32] | LEE J H, YADOLLAHPOUR P, WATKINS A, et al.. EquiFold: protein structure prediction with a novel coarse-grained structure representation[J/OL]. bioRxiv, 2022[2025-10-21]. . |
| [33] | EVANS R, O'NEILL M, PRITZEL A, et al.. Protein complex prediction with AlphaFold-multimer[J/OL]. bioRxiv, 2021: 2015-2021[2025-10-21]. . |
| [34] | ABRAMSON J, ADLER J, DUNGER J, et al.. Accurate structure prediction of biomolecular interactions with AlphaFold 3[J]. Nature, 2024, 630(8016): 493-500. |
| [35] | JUMPER J, EVANS R, PRITZEL A, et al.. Highly accurate protein structure prediction with AlphaFold[J]. Nature, 2021, 596(7873): 583-589. |
| [36] | HAYES T, RAO R, AKIN H, et al.. Simulating 500 million years of evolution with a language model[J]. Science, 2025, 387(6736): 850-858. |
| [37] | JOUBBI S, MACCARI G, CIANO G, et al.. Improving antibody-antigen interaction prediction through flexibility with ESMFold[C]// Proceedings of the 18th International Joint Conference on Biomedical Engineering Systems and Technologies. SCITEPRESS-Science and Technology Publications, 2025: 603-610. |
| [38] | JOUBBI S, MICHELI A, MILAZZO P, et al.. Antibody design using deep learning: from sequence and structure design to affinity maturation[J/OL]. Brief. Bioinform., 2024, 25(4): bbae307[2025-09-30]. . |
| [39] | LI M, KANG L, XIONG Y, et al.. SESNet: sequence-structure feature-integrated deep learning method for data-efficient protein engineering[J/OL]. J. Cheminf., 2023, 15(1): 12[2025-12-21]. . |
| [40] | SHIN J E, RIESSELMAN A J, KOLLASCH A W, et al.. Protein design and variant prediction using autoregressive generative models[J/OL]. Nat. Commun., 2021, 12(1): 2403[2025-12-21]. . |
| [41] | HU R, FU L, CHEN Y, et al.. Protein engineering via Bayesian optimization-guided evolutionary algorithm and robotic experiments[J/OL]. Brief. Bioinform., 2023, 24(1): bbac570[2025-09-30]. . |
| [42] | KHAN A, COWEN-RIVERS A I, GROSNIT A, et al.. Toward real-world automated antibody design with combinatorial Bayesian optimization[J/OL]. Cell Rep. Methods, 2023, 3(1): 100374[2025-09-30]. . |
| [43] | FERRUZ N, SCHMIDT S, HÖCKER B. ProtGPT2 is a deep unsupervised language model for protein design[J/OL]. Nat. Commun., 2022, 13(1): 4348[2025-12-21]. . |
| [44] | CHUNGYOUN M, RUFFOLO J, GRAY J. FLAb: benchmarking deep learning methods for antibody fitness prediction[J/OL]. bioRxiv, 2024: 2021-2024[2025-10-21]. . |
| [45] | LIAO Y, MA H, WANG Z, et al.. Rapid restoration of potent neutralization activity against the latest Omicron variant JN.1 via AI rational design and antibody engineering[J/OL]. Proc. Natl. Acad. Sci. USA, 2025, 122(6): e2406659122[2025-09-30]. . |
| [46] | GUO P, LI M, PAN H, et al.. Multi-modality representation learning for antibody-antigen interactions prediction[C]// 2025 IEEE International Conference on Multimedia and Expo (ICME). Piscataway, New Jersey: IEEE, 2025: 1-6. |
| [47] | HOSSAIN D, SAGHAPOUR E, SONG K, et al.. Llama-affinity: a predictive antibody antigen binding model integrating antibody sequences with Llama3 backbone architecture[EB/OL]. arXiv, 2025: 2506.09052[2025-09-30]. . |
| [48] | AGARWAL A A, HARRANG J, NOBLE D, et al.. AlphaBind, a domain-specific model to predict and optimize antibody-antigen binding affinity[J/OL]. MAbs, 2025, 17(1): 2534626[2025-12-21]. . |
| [49] | ROLLINS Z A, WIDATALLA T, CHENG A C, et al.. AbMelt: learning antibody thermostability from molecular dynamics[J]. Biophys. J., 2024, 123(17): 2921-2933. |
| [50] | MAKOWSKI E K, KINNUNEN P C, HUANG J, et al.. Co-optimization of therapeutic antibody affinity and specificity using machine learning models that generalize to novel mutational space[J/OL]. Nat. Commun., 2022, 13(1): 3788[2025-09-30]. . |
| [51] | ZHOU X, XUE D, CHEN R, et al.. Antigen-specific antibody design via direct energy-based preference optimization[EB/OL]. arXiv, 2024: 2403.16576[2025-09-30]. . |
| [1] | 李明, 曲楠楠, 李伟. 抗体药物连续生物工艺的技术突破与监管挑战[J]. 生物技术进展, 2025, 15(5): 845-853. |
| [2] | 杨懿祺, 张志高, 游小龙, 张婧, 林冠峰, 吴英松. 抗体药物的发展与应用[J]. 生物技术进展, 2022, 12(3): 358-365. |
| [3] | 辛志奇, 赵航, 汪海, 路铁刚. 基于深度学习的作物基因组学和遗传改良[J]. 生物技术进展, 2021, 11(4): 483-488. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||