


生物技术进展 ›› 2025, Vol. 15 ›› Issue (6): 1020-1030.DOI: 10.19586/j.2095-2341.2025.0085
• 研究论文 • 上一篇
乔华亮1,2( ), 王娇1, 焦博1, 柳洪2, 李俊明2, 王耕2, 周春江2, 周硕1(
), 王娇1, 焦博1, 柳洪2, 李俊明2, 王耕2, 周春江2, 周硕1( )
)
收稿日期:2025-07-28
接受日期:2025-10-22
出版日期:2025-11-25
发布日期:2026-01-04
通讯作者:
周硕
作者简介:乔华亮E-mail: qiaohl1006@163.com;
基金资助:
Hualiang QIAO1,2( ), Jiao WANG1, Bo JIAO1, Hong LIU2, Junming LI2, Geng WANG2, Chunjiang ZHOU2, Shuo ZHOU1(
), Jiao WANG1, Bo JIAO1, Hong LIU2, Junming LI2, Geng WANG2, Chunjiang ZHOU2, Shuo ZHOU1( )
)
Received:2025-07-28
Accepted:2025-10-22
Online:2025-11-25
Published:2026-01-04
Contact:
Shuo ZHOU
摘要:
小麦TaWRKY13转录因子是调控叶片衰老的重要基因,对其作用机制进行解析可影响农作物的产量和品质。利用农杆菌转化法获得了TaWRKY13的过表达小麦,并观察到TaWRKY13能够促进叶片衰老。通过外源脱落酸(abscisic acid,ABA)处理,发现TaWRKY13过表达株系能够响应ABA的处理,并可加速TaWRKY13过表达衰老进程。通过实时定量PCR、ChIP-PCR和凝胶迁移(electrophoretic mobility shift assay,EMSA)实验证明TaWRKY13能够通过直接结合TaNCED5影响ABA的变化,进而改变叶片衰老进程。综上,TaWRKY13直接结合TaNCED5调控ABA的合成,影响ABA的含量,进而影响叶片衰老的进程。结果可为阐明小麦中叶片的衰老机制提供理论依据。
中图分类号:
乔华亮, 王娇, 焦博, 柳洪, 李俊明, 王耕, 周春江, 周硕. 小麦TaWRKY13转录因子参与ABA介导叶片衰老的功能研究[J]. 生物技术进展, 2025, 15(6): 1020-1030.
Hualiang QIAO, Jiao WANG, Bo JIAO, Hong LIU, Junming LI, Geng WANG, Chunjiang ZHOU, Shuo ZHOU. Functional Analysis of the Wheat TaWRKY13 Transcription Factor in ABA-mediated Leaf Senescence[J]. Current Biotechnology, 2025, 15(6): 1020-1030.
| 用途 | 载体 | 细菌内抗生素 | 植物内抗生菌 |
|---|---|---|---|
| 原核表达载体 | pMal-C2X | 氨苄(ampicillin, | 利福平(rifampicin, Rfp) |
| 过表达植物载体 | pTCK303 | 卡那(kanamycin, Kan) | 潮霉素(hygromycin, Hyg) |
表1 本文所涉及的载体
Table 1 The carrier involved in this article
| 用途 | 载体 | 细菌内抗生素 | 植物内抗生菌 |
|---|---|---|---|
| 原核表达载体 | pMal-C2X | 氨苄(ampicillin, | 利福平(rifampicin, Rfp) |
| 过表达植物载体 | pTCK303 | 卡那(kanamycin, Kan) | 潮霉素(hygromycin, Hyg) |
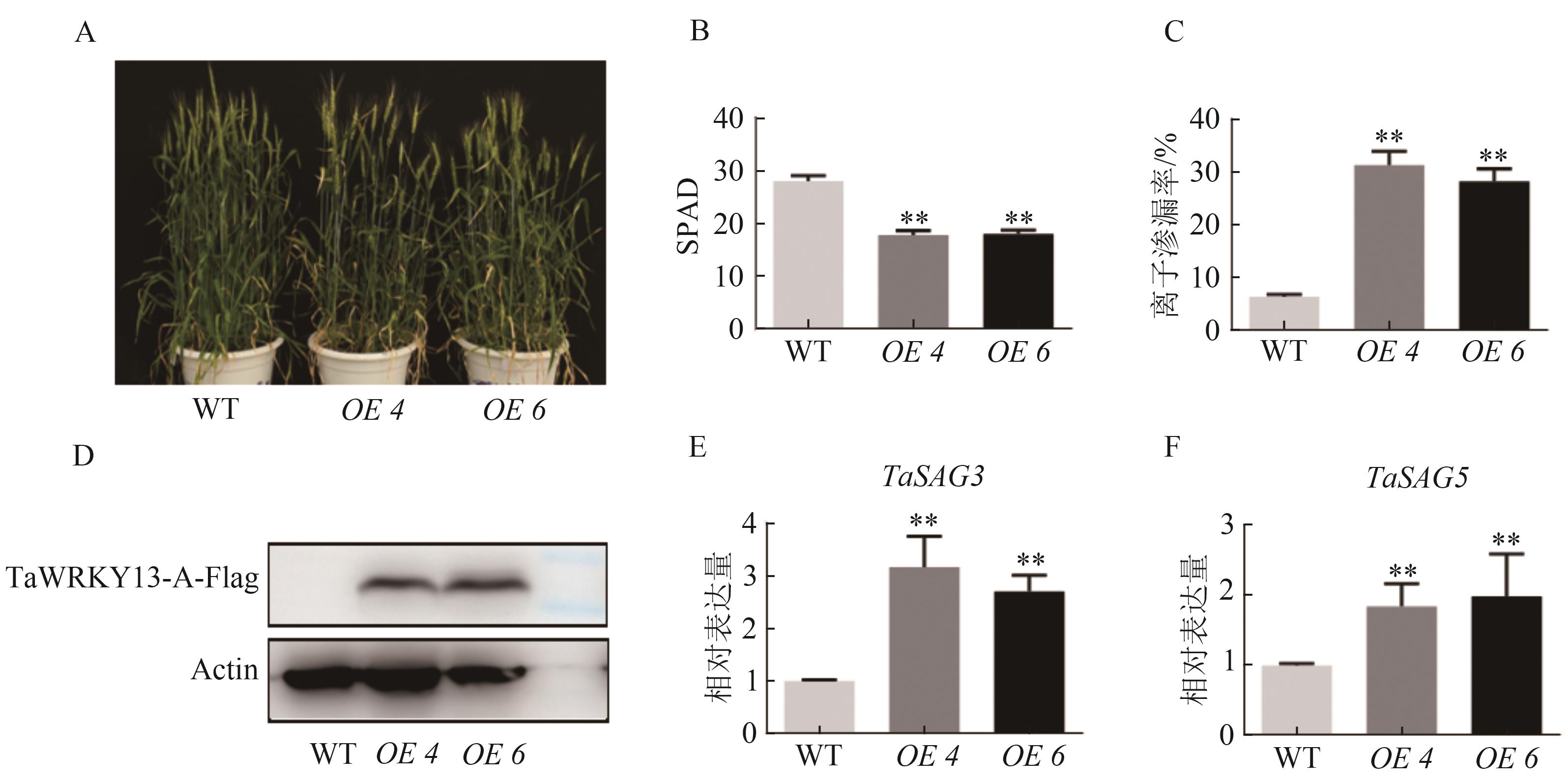
图2 TaWRKY13过表达小麦表型观察A:TaWRKY13转基因小麦表型;B:叶绿素含量;C:离子渗漏率;D:蛋白表达量;E~F:叶片衰老maker基因(TaSAG3、TaSAG5)表达量。**表示与WT相比在P0.01水平上有统计学差异。
Fig. 2 Observation of phenotype of TaWRKY13 overexpression in wheat
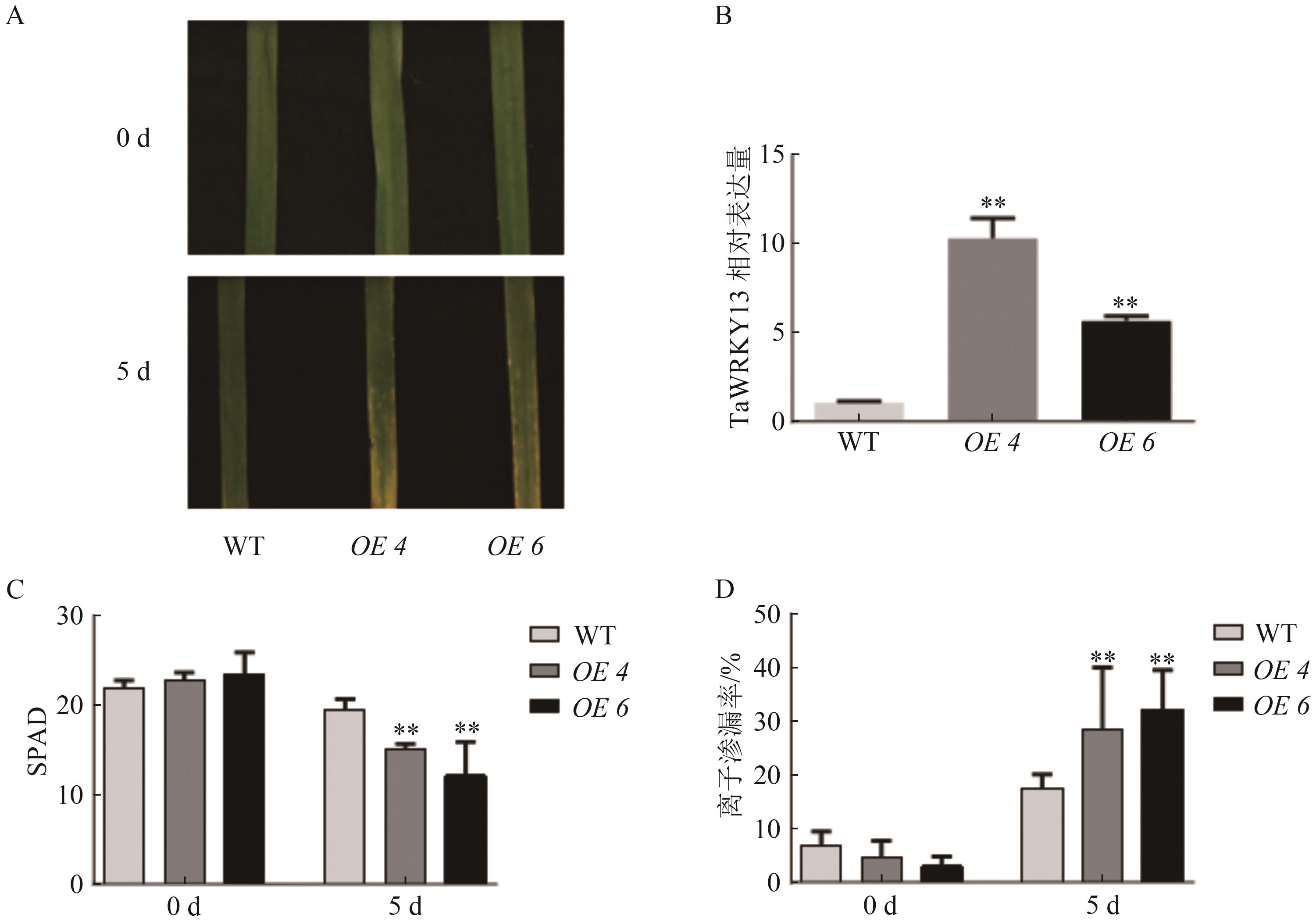
图3 TaWRKY13过表达小麦叶片在黑暗处理下表型A:黑暗处理下过表达小麦叶片表型;B:TaWRKY13过表达小麦转录水平表达检测;C:叶绿素含量;D:离子渗漏率。**表示与WT相比在P0.01水平上有统计学差异。
Fig. 3 Phenotype of wheat leaves overexpressing TaWRKY13 under dark treatment
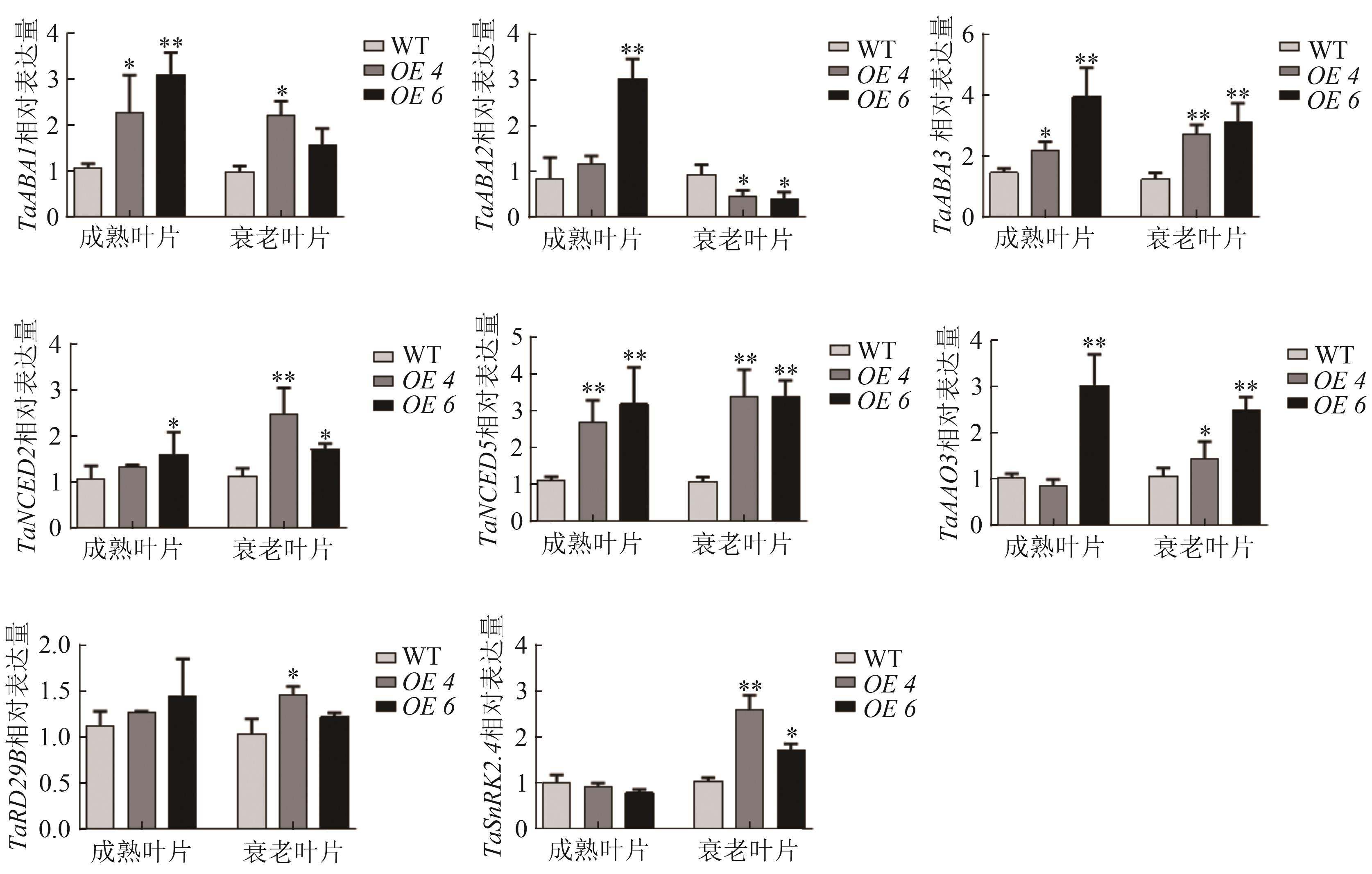
图6 TaWRKY13过表达小麦中ABA合成和信号通路相关基因检测注:*,**分别表示与WT组相比在P0.05和P0.01水平上有统计学差异。
Fig. 6 Detection of ABA synthesis and signaling pathway-related genes in wheat overexpressing TaWRKY13
| [1] | TANAKA A, ITO H. Chlorophyll degradation and its physiological function[J]. Plant Cell Physiol., 2025, 66(2): 139-152. |
| [2] | LI Y, CAO T, GUO Y, et al.. Regulatory and retrograde signaling networks in the chlorophyll biosynthetic pathway[J]. J. Integr. Plant Biol., 2025, 67(4): 887-911. |
| [3] | WOJCIECHOWSKA N, SOBIESZCZUK-NOWICKA E, BAGNIEWSKA-ZADWORNA A. Plant organ senescence-regulation by manifold pathways[J]. Plant Biol., 2018, 20(2): 167-181. |
| [4] | GUO Y, REN G, ZHANG K, et al.. Leaf senescence: progression, regulation, and application[J/OL]. Mol. Hortic., 2021, 1(1): 5[2025-09-17]. . |
| [5] | LIM P O, KIM H J, NAM H G. Leaf senescence[J]. Annu. Rev. Plant Biol., 2007, 58: 115-136. |
| [6] | YANG H, LI Y, QIAO Y, et al.. Low light stress promotes new tiller regeneration by changing source-sink relationship and activating expression of expansin genes in wheat[J]. Plant Cell Environ., 2023, 46(5): 1562-1581. |
| [7] | MA B, ZHANG L, HE Z. Understanding the regulation of cereal grain filling: the way forward[J]. J. Integr. Plant Biol., 2023, 65(2): 526-547. |
| [8] | WOO H R, KIM H J, LIM P O, et al.. Leaf senescence: systems and dynamics aspects[J]. Annu. Rev. Plant Biol., 2019, 70: 347-376. |
| [9] | ZHAO Y, LIU H, CAO J, et al.. FORKHEAD BOX1 promotes leaf senescence via the histone acetyltransferase HAC1 and the transcription factors TGA7 and ABF2/3[J/OL]. Plant Cell, 2025, 37(7): koaf170[2025-09-17]. . |
| [10] | GUO X, WANG Y, ZHAO C, et al.. An Arabidopsis single-nucleus atlas Decodes leaf senescence and nutrient allocation[J]. Cell, 2025, 188(11): 2856-2871. |
| [11] | ZHANG H, ZHOU C. Signal transduction in leaf senescence[J]. Plant Mol. Biol., 2013, 82(6): 539-545. |
| [12] | HUANG P, LI Z, GUO H. New advances in the regulation of leaf senescence by classical and peptide hormones[J/OL]. Front. Plant Sci., 2022, 13: 923136[2025-09-17]. . |
| [13] | WAADT R, SELLER C A, HSU P K, et al.. Plant hormone regulation of abiotic stress responses[J]. Nat. Rev. Mol. Cell Biol., 2022, 23(10): 680-694. |
| [14] | ALI J, MUKARRAM M, OJO J, et al.. Harnessing phytohormones: advancing plant growth and defence strategies for sustainable agriculture[J/OL]. Physiol. Plant., 2024, 176(3): e14307[2025-09-17]. . |
| [15] | TANG X, MEI Y, HE K, et al.. The RING-type E3 ligase RIE1 sustains leaf longevity by specifically targeting AtACS7 to fine-tune ethylene production in Arabidopsis [J/OL]. Proc. Natl. Acad. Sci. USA, 2024, 121(48): e2411271121[2025-09-17]. . |
| [16] | LIM C, KANG K, LIM J, et al.. RICE LONG GRAIN 3 delays dark-induced senescence by downregulating abscisic acid signaling and upregulating reactive oxygen species scavenging activity[J]. Plant J., 2024, 120(4): 1474-1487. |
| [17] | ZHANG Y, TAN S, GAO Y, et al.. CLE42 delays leaf senescence by antagonizing ethylene pathway in Arabidopsis [J]. New Phytol., 2022, 235(2): 550-562. |
| [18] | FANG C, ZHANG H, WAN J, et al.. Control of leaf senescence by an MeOH-jasmonates cascade that is epigenetically regulated by OsSRT1 in rice[J]. Mol. Plant, 2016, 9(10): 1366-1378. |
| [19] | XIE W, XUE X, WANG Y, et al.. Natural mutation in Stay-Green (OsSGR) confers enhanced resistance to rice sheath blight through elevating cytokinin content[J]. Plant Biotechnol. J., 2025, 23(3): 807-823. |
| [20] | CHEN K, LI G J, BRESSAN R A, et al.. Abscisic acid dynamics, signaling, and functions in plants[J]. J. Integr. Plant Biol., 2020, 621: 25-54. |
| [21] | LI Y, KAMIYAMA Y, MINEGISHI F, et al.. Group C MAP kinases phosphorylate MBD10 to regulate ABA-induced leaf senescence in Arabidopsis [J]. Plant J., 2024, 118(6): 1747-1759. |
| [22] | YANG F, ZHAO L L, SONG L Q, et al.. Apple E3 ligase MdPUB23 mediates ubiquitin-dependent degradation of MdABI5 to delay ABA-triggered leaf senescence[J/OL]. Hortic. Res., 2024, 11(4): uhae029[2025-09-17]. . |
| [23] | ASAD M A U, ZAKARI S A, ZHAO Q, et al.. Abiotic stresses intervene with ABA signaling to induce destructive metabolic pathways leading to death: premature leaf senescence in plants[J/OL]. Int. J. Mol. Sci., 2019, 20(2): 256[2025-09-17]. . |
| [24] | ZHAO Y, CHAN Z, GAO J, et al.. ABA receptor PYL9 promotes drought resistance and leaf senescence[J]. Proc. Natl. Acad. Sci. USA, 2016, 113(7): 1949-1954. |
| [25] | WANG G, LIU X, GAN S S. The ABA-AtNAP-SAG113 PP2C module regulates leaf senescence by dephoshorylating SAG114 SnRK3.25 in Arabidopsis [J/OL]. Mol. Hortic., 2023, 3(1): 22[2025-09-17]. . |
| [26] | PANOZZO A, BOLLA P K, BARION G, et al.. Phytohormonal regulation of abiotic stress tolerance, leaf senescence and yield response in field crops: a comprehensive review[J/OL]. BioTech, 2025, 14(1): 14[2025-09-17]. . |
| [27] | CAO J, LIU H, TAN S, et al.. Transcription factors-regulated leaf senescence: current knowledge, challenges and approaches[J/OL]. Int. J. Mol. Sci., 2023, 24(11): 9245[2025-09-17]. . |
| [28] | MAO C, LU S, LV B, et al.. A rice NAC transcription factor promotes leaf senescence via ABA biosynthesis[J]. Plant Physiol., 2017, 174(3): 1747-1763. |
| [29] | WEN B, ZHAO X, GONG X, et al.. The NAC transcription factor MdNAC4 positively regulates nitrogen deficiency-induced leaf senescence by enhancing ABA biosynthesis in apple[J/OL]. Mol. Hortic., 2023, 3(1): 5[2025-09-17]. . |
| [30] | MA L, GAO Q, LIU Y, et al.. NAP-dependent crosstalk between ethylene biosynthesis and abscisic acid signaling pathway coordinately modulates leaf senescence in plants[J/OL]. Plant J., 2025, 122(5): e70245[2025-09-17]. . |
| [31] | MENG L, YANG H, YANG J, et al.. Tulip transcription factor TgWRKY75 activates salicylic acid and abscisic acid biosynthesis to synergistically promote petal senescence[J]. J. Exp. Bot., 2024, 75(8): 2435-2450. |
| [32] | LEHTI-SHIU M D, PANCHY N, WANG P, et al.. Diversity, expansion, and evolutionary novelty of plant DNA-binding transcription factor families[J]. Biochim. Biophys. Acta Gene Regul. Mech., 2017, 1860(1): 3-20. |
| [33] | STRADER L, WEIJERS D, WAGNER D. Plant transcription factors: being in the right place with the right company[J/OL]. Curr. Opin. Plant Biol., 2022, 65: 102136[2025-09-17]. . |
| [34] | WANG Y, LU W, DENG D. Bioinformatic landscapes for plant transcription factor system research[J]. Planta, 2016, 243(2): 297-304. |
| [35] | ZHANG Y, LU Y, SAYYED H E L, et al.. Transcription factor dynamics in plants: insights and technologies for in vivo imaging[J]. Plant Physiol., 2022, 189(1): 23-36. |
| [36] | XIONG H, HE H, CHANG Y, et al.. Multiple roles of NAC transcription factors in plant development and stress responses[J]. J. Integr. Plant Biol., 2025, 67(3): 510-538. |
| [37] | AHMAD Z, RAMAKRISHNAN M, WANG C, et al.. Unravelling the role of WRKY transcription factors in leaf senescence: genetic and molecular insights[J]. J. Adv. Res., 2025, 74: 191-206. |
| [38] | NG D W, ABEYSINGHE J K, KAMALI M. Regulating the regulators: the control of transcription factors in plant defense signaling[J/OL]. Int. J. Mol. Sci., 2018, 19(12): 3737[2025-09-17]. . |
| [39] | 于延冲,乔孟,刘振华,等.WRKY转录因子功能的多样化[J].生命科学, 2010, 22(4): 345-351. |
| YU Y C, QIAO M, LIU Z H, et al.. Diversification of WRKY transcription factor[J]. Bull. Life Sci., 2010, 22(4): 345-351. | |
| [40] | 郝中娜,陶荣祥.WRKY转录因子超家族的研究[J].生命科学,2006(2):175-179. |
| HAO Z N, TAO R X. Study of WRKY transcription factor superfamily[J]. Chin. Bull. Life Sci., 2006(2): 175-179. | |
| [41] | YANG L, FANG S, LIU L, et al.. WRKY transcription factors: Hubs for regulating plant growth and stress responses[J]. J. Integr. Plant Biol., 2025, 67(3): 488-509. |
| [42] | RUSHTON P J, SOMSSICH I E, RINGLER P, et al.. WRKY transcription factors[J]. Trends Plant Sci., 2010, 15(5): 247-258. |
| [43] | WANI S H, ANAND S, SINGH B, et al.. WRKY transcription factors and plant defense responses: latest discoveries and future prospects[J]. Plant Cell Rep., 2021, 40(7): 1071-1085. |
| [44] | QIAO H, LIU Y, CHENG L, et al.. TaWRKY13-a serves as a mediator of jasmonic acid-related leaf senescence by modulating jasmonic acid biosynthesis[J/OL]. Front. Plant Sci., 2021, 12: 717233[2025-09-17]. . |
| [45] | ZHAO M M, ZHANG X W, LIU Y W, et al.. A WRKY transcription factor, TaWRKY42-B, facilitates initiation of leaf senescence by promoting jasmonic acid biosynthesis[J/OL]. BMC Plant Biol., 2020, 20(1): 444[2025-09-17]. . |
| [46] | YU Y, QI Y, XU J, et al.. Arabidopsis WRKY71 regulates ethylene-mediated leaf senescence by directly activating EIN2 ORE1 and ACS2 genes[J]. Plant J., 2021, 107(6): 1819-1836. |
| [47] | KIM T, KANG K, KIM S H, et al.. OsWRKY5 promotes rice leaf senescence via senescence-associated NAC and abscisic acid biosynthesis pathway[J/OL]. Int. J. Mol. Sci., 2019, 20(18): 4437[2025-09-17]. . |
| [48] | ZENTGRAF U, DOLL J. Arabidopsis WRKY53, a node of multi-layer regulation in the network of senescence[J/OL]. Plants, 2019, 8(12): 578[2025-09-17]. . |
| [49] | MIAO Y, ZENTGRAF U. The antagonist function of Arabidopsis WRKY53 and ESR/ESP in leaf senescence is modulated by the jasmonic and salicylic acid equilibrium[J]. Plant Cell, 2007, 19(3): 819-830. |
| [50] | MA Q, XU S, HU S, et al.. WRKY75-mediated transcriptional regulation of OASA1 controls leaf senescence in Arabidopsis [J/OL]. Physiol. Plant., 2025, 177(2): e70193[2025-09-17]. . |
| [51] | ZHANG W, TANG S, LI X, et al.. Arabidopsis WRKY1 promotes monocarpic senescence by integrative regulation of flowering, leaf senescence, and nitrogen remobilization[J]. Mol. Plant, 2024, 17(8): 1289-1306. |
| [52] | ZHAO L, ZHANG W, SONG Q, et al.. A WRKY transcription factor, TaWRKY40-D, promotes leaf senescence associated with jasmonic acid and abscisic acid pathways in wheat[J]. Plant Biol., 2020, 22(6): 1072-1085. |
| [53] | ZHOU S, ZHENG W J, LIU B H, et al.. Characterizing the role of TaWRKY13 in salt tolerance[J/OL]. Int. J. Mol. Sci., 2019, 20(22): 5712[2025-09-17]. . |
| [54] | MA J, GAO X, LIU Q, et al.. Overexpression of TaWRKY146 increases drought tolerance through inducing stomatal closure in Arabidopsis thaliana [J/OL]. Front. Plant Sci., 2017, 8: 2036[2025-09-17]. . |
| [55] | LIU X, KIM Y J, MÜLLER R, et al.. AGAMOUS terminates floral stem cell maintenance in Arabidopsis by directly repressing WUSCHEL through recruitment of Polycomb Group proteins[J]. Plant Cell, 2011, 23(10): 3654-3670. |
| [56] | JIANG J, MA S, YE N, et al.. WRKY transcription factors in plant responses to stresses[J]. J. Integr. Plant Biol., 2017, 59(2): 86-101. |
| [57] | NAN H, GAO L Z. Genome-wide analysis of WRKY genes and their response to hormone and mechanic stresses in carrot[J/OL]. Front. Genet., 2019, 10: 363[2025-09-17]. . |
| [58] | GOYAL P, MANZOOR M M, VISHWAKARMA R A, et al.. A comprehensive transcriptome-wide identification and screening of WRKY gene family engaged in abiotic stress in Glycyrrhiza glabra [J/OL]. Sci. Rep., 2020, 10(1): 373[2025-09-17]. . |
| [59] | KANG G, YAN D, CHEN X, et al.. HbWRKY82, a novel Ⅱc WRKY transcription factor from Hevea brasiliensis associated with abiotic stress tolerance and leaf senescence in Arabidopsis [J]. Physiol. Plant., 2021, 171(1): 151-160. |
| [60] | ANDRADE GALAN A G, DOLL J, VON ROEPENACK-LAHAYE E, et al.. The transcription factor WRKY25 can act as redox switch to drive the expression of WRKY53 during leaf senescence in Arabidopsis [J/OL]. Sci. Rep., 2025, 15(1): 27623[2025-09-17]. . |
| [61] | TAN Q, ZHAO M, GAO J, et al.. AtVQ25 promotes salicylic acid-related leaf senescence by fine-tuning the self-repression of AtWRKY53[J]. J. Integr. Plant Biol., 2024, 66(6): 1126-1147. |
| [62] | YU J, ZHU C, XUAN W, et al.. Genome-wide association studies identify OsWRKY53 as a key regulator of salt tolerance in rice[J/OL]. Nat. Commun., 2023, 14(1): 3550[2025-09-17]. . |
| [63] | MENG Y, LV Q, LI L, et al.. E3 ubiquitin ligase TaSDIR1-4A activates membrane-bound transcription factor TaWRKY29 to positively regulate drought resistance[J]. Plant Biotechnol. J., 2024, 22(4): 987-1000. |
| [64] | JIANG Y, LIANG G, YANG S, et al.. Arabidopsis WRKY57 functions as a node of convergence for jasmonic acid- and auxin-mediated signaling in jasmonic acid-induced leaf senescence[J]. Plant Cell, 2014, 26(1): 230-245. |
| [65] | BESSEAU S, LI J, PALVA E T. WRKY54 and WRKY70 co-operate as negative regulators of leaf senescence in Arabidopsis thaliana [J]. J. Exp. Bot., 2012, 63(7): 2667-2679. |
| [66] | ZHANG H, ZHANG L, WU S, et al.. AtWRKY75 positively regulates age-triggered leaf senescence through gibberellin pathway[J]. Plant Divers., 2021, 43(4): 331-340. |
| [67] | CHEN L, XIANG S, CHEN Y, et al.. Arabidopsis WRKY45 interacts with the DELLA protein RGL1 to positively regulate age-triggered leaf senescence[J]. Mol. Plant, 2017, 10(9): 1174-1189. |
| [68] | ZHENG Y, GE J, BAO C, et al.. Histone deacetylase HDA9 and WRKY53 transcription factor are mutual antagonists in regulation of plant stress response[J]. Mol. Plant, 2020, 13(4): 598-611. |
| [69] | WANG D, WEI L, MA J, et al.. Bacillus cereus NJ01 induces SA- and ABA-mediated immunity against bacterial pathogens through the EDS1-WRKY18 module[J/OL]. Cell Rep., 2024, 43(4): 113985[2025-09-17]. . |
| [70] | FREY A, EFFROY D, LEFEBVRE V, et al.. Epoxycarotenoid cleavage by NCED5 fine-tunes ABA accumulation and affects seed dormancy and drought tolerance with other NCED family members[J]. Plant J., 2012, 70(3): 501-512. |
| [1] | 陈中惠, 李景蕻. 外源植物激素对高温胁迫下水培绿萝耐热性的影响[J]. 生物技术进展, 2023, 13(5): 742-747. |
| [2] | 戚义东,秦华,高雅迪,王芳芳,黄荣峰,权瑞党. 脱落酸拮抗赤霉素抑制水稻地上部生长的研究[J]. 生物技术进展, 2019, 9(5): 483-489. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||