


生物技术进展 ›› 2025, Vol. 15 ›› Issue (6): 1003-1019.DOI: 10.19586/j.2095-2341.2025.0145
• 研究论文 •
张凯鹏1,2,3( ), 王莉4, 唐莉4, 李婧1,2,3, 陈琪4(
), 王莉4, 唐莉4, 李婧1,2,3, 陈琪4( )
)
收稿日期:2025-10-21
接受日期:2025-11-12
出版日期:2025-11-25
发布日期:2026-01-04
通讯作者:
陈琪
作者简介:张凯鹏E-mail: zkpeng0626@163.com;
基金资助:
Kaipeng ZHANG1,2,3( ), Li WANG4, Li TANG4, Jing LI1,2,3, Qi CHEN4(
), Li WANG4, Li TANG4, Jing LI1,2,3, Qi CHEN4( )
)
Received:2025-10-21
Accepted:2025-11-12
Online:2025-11-25
Published:2026-01-04
Contact:
Qi CHEN
摘要:
研究旨在对赤松梢斑螟(Dioryctria sylvestrella)气味降解酶(odorant-degrading enzyme, ODE)相关基因进行鉴定、分类及组织表达模式分析,以期为该害虫的绿色高效防治技术提供理论依据。在赤松梢斑螟雌、雄触角转录组数据中筛选出4种ODEs基因:羧酸酯酶(carboxylesterases, CXEs)、醛氧化酶(aldehyde oxidases, AOXs)、谷胱甘肽转移酶(glutathione S-transferases, GSTs)和细胞色素P450氧化酶(cytochrome P450s, CYPs),对这些基因进行生物信息学分析和同源建树、命名;选取触角中FPKM值较高的候选基因进行后续验证,并采用RT-qPCR技术分析所选基因在成虫不同组织中的特异表达情况。研究共鉴定出137个候选ODEs相关基因,包括39个DsylCXEs、6个DsylAOXs、32个DsylGSTs和60个DsylCYPs。RT-qPCR结果表明,18个DsylODEs在触角中的表达量显著高于其他组织。在赤松梢斑螟的触角中显著表达的ODEs基因类型主要为谷胱甘肽转移酶与细胞色素P450。研究首次筛选出了潜在的具有较强气味降解能力的赤松梢斑螟ODEs基因,为进一步探究赤松梢斑螟在嗅觉识别过程中的气味降解机制奠定了基础。
中图分类号:
张凯鹏, 王莉, 唐莉, 李婧, 陈琪. 赤松梢斑螟气味降解酶基因筛选及其在不同组织的表达差异研究[J]. 生物技术进展, 2025, 15(6): 1003-1019.
Kaipeng ZHANG, Li WANG, Li TANG, Jing LI, Qi CHEN. Study on Screening of Odorant Degradation Enzyme Genes in Dioryctria sylvestrella and its Expression Level in Different Tissues[J]. Current Biotechnology, 2025, 15(6): 1003-1019.
| 基因名称 | 正向引物(5′→3′) | 反向引物(5′→3′) |
|---|---|---|
| DsylCXE17 | 5′-CTTGGCCATGGTAAAGAAGC-3′ | 5′-TGAGAGGGTCTGGTCTTGCT-3′ |
| DsylCXE20 | 5′-AGGAAAACTGCAAGGCAAGA-3′ | 5′-CAAATCTCAGCTCACCGACA-3′ |
| DsylCXE30 | 5′-ATCCATGGAGGAGGCTTTTT-3′ | 5′-AGAGAAATCCGAGTGCCTCA-3′ |
| DsylCXE32 | 5′-ATCACGGGTAGCGAAGATTG-3′ | 5′-TGCACCACCGTGAATAAAGA-3′ |
| DsylCXE38 | 5′-CCTTCCGCTCTGTCGTTATC-3′ | 5′-GAGGAGTTGTTCAGGCTTGC-3′ |
| DsylCXE39 | 5′-ATCTGGCACCAGGCAGTATC-3′ | 5′-ACATCTCCTCCCACACCTTG-3′ |
| DsylAOX2 | 5′-CTCAAAGACTGTGGGGGAAA-3′ | 5′-TTGAGTTGTGGATGCTCTGC-3′ |
| DsylAOX4 | 5′-ACCCGTACGCTGTAATCGAC-3′ | 5′-CTGCGGATAGCTGGAGAAAG-3′ |
| DsylGST10 | 5′-GCCTTCGGCTATGACTTCAG-3′ | 5′-ATTTGTCCTCCTGCATGGTC-3′ |
| DsylGST14 | 5′-GACGCCGGAATATTTGAAGA-3′ | 5′-TTCGGGTACAGGGAGTCATC-3′ |
| DsylGST15 | 5′-AAATCCCCAGCATACAGTGC-3′ | 5′-TCTCTTCAGGGGGTCATTTG-3′ |
| DsylGST20 | 5′-GGAAGATGAGGCTGTCAAGG-3′ | 5′-AGCAGCTAGGTGTCCGTTGT-3′ |
| DsylGST23 | 5′-GGAAGATGAGGCTGTCAAGG-3′ | 5′-AGCAGCTAGGTGTCCGTTGT-3′ |
| DsylGST30 | 5′-GGAAGATGAGGCTGTCAAGG-3′ | 5′-AGCAGCTAGGTGTCCGTTGT-3′ |
| DsylCYP19 | 5′-CCGTGTGGGAGAAATGACTT-3′ | 5′-CATCACGATGTTTTCCGATG-3′ |
| DsylCYP30 | 5′-GGATGGCTCGGTCACATACT-3′ | 5′-CTGCCGTAGCCAATGATTTT-3′ |
| DsylCYP32 | 5′-AGGCCGAACATAACCACAAG-3′ | 5′-CGTTGAATTGCGAAGTTTGA-3′ |
| DsylCYP36 | 5′-ACGCTAACATCTGGGGTCAC-3′ | 5′-ACCCCAATACTTCGGGTTTC-3′ |
| DsylCYP37 | 5′-ATGCGTTTGTACCCTTCTCG-3′ | 5′-TCGGAAATTTCTCAGGATGG-3′ |
| DsylCYP55 | 5′-CCCTGTGGGGCTGTTATAGA-3′ | 5′-CCGGAAAGAAATCTGGATGA-3′ |
| DsylGAPDH | 5′-TGCACAACTAACTGCCTTGC-3′ | 5′-TGGAAGCGGGAATTATGTTC-3′ |
表 1 实时荧光定量PCR所用引物
Table 1 The primers used for the RT-qPCR
| 基因名称 | 正向引物(5′→3′) | 反向引物(5′→3′) |
|---|---|---|
| DsylCXE17 | 5′-CTTGGCCATGGTAAAGAAGC-3′ | 5′-TGAGAGGGTCTGGTCTTGCT-3′ |
| DsylCXE20 | 5′-AGGAAAACTGCAAGGCAAGA-3′ | 5′-CAAATCTCAGCTCACCGACA-3′ |
| DsylCXE30 | 5′-ATCCATGGAGGAGGCTTTTT-3′ | 5′-AGAGAAATCCGAGTGCCTCA-3′ |
| DsylCXE32 | 5′-ATCACGGGTAGCGAAGATTG-3′ | 5′-TGCACCACCGTGAATAAAGA-3′ |
| DsylCXE38 | 5′-CCTTCCGCTCTGTCGTTATC-3′ | 5′-GAGGAGTTGTTCAGGCTTGC-3′ |
| DsylCXE39 | 5′-ATCTGGCACCAGGCAGTATC-3′ | 5′-ACATCTCCTCCCACACCTTG-3′ |
| DsylAOX2 | 5′-CTCAAAGACTGTGGGGGAAA-3′ | 5′-TTGAGTTGTGGATGCTCTGC-3′ |
| DsylAOX4 | 5′-ACCCGTACGCTGTAATCGAC-3′ | 5′-CTGCGGATAGCTGGAGAAAG-3′ |
| DsylGST10 | 5′-GCCTTCGGCTATGACTTCAG-3′ | 5′-ATTTGTCCTCCTGCATGGTC-3′ |
| DsylGST14 | 5′-GACGCCGGAATATTTGAAGA-3′ | 5′-TTCGGGTACAGGGAGTCATC-3′ |
| DsylGST15 | 5′-AAATCCCCAGCATACAGTGC-3′ | 5′-TCTCTTCAGGGGGTCATTTG-3′ |
| DsylGST20 | 5′-GGAAGATGAGGCTGTCAAGG-3′ | 5′-AGCAGCTAGGTGTCCGTTGT-3′ |
| DsylGST23 | 5′-GGAAGATGAGGCTGTCAAGG-3′ | 5′-AGCAGCTAGGTGTCCGTTGT-3′ |
| DsylGST30 | 5′-GGAAGATGAGGCTGTCAAGG-3′ | 5′-AGCAGCTAGGTGTCCGTTGT-3′ |
| DsylCYP19 | 5′-CCGTGTGGGAGAAATGACTT-3′ | 5′-CATCACGATGTTTTCCGATG-3′ |
| DsylCYP30 | 5′-GGATGGCTCGGTCACATACT-3′ | 5′-CTGCCGTAGCCAATGATTTT-3′ |
| DsylCYP32 | 5′-AGGCCGAACATAACCACAAG-3′ | 5′-CGTTGAATTGCGAAGTTTGA-3′ |
| DsylCYP36 | 5′-ACGCTAACATCTGGGGTCAC-3′ | 5′-ACCCCAATACTTCGGGTTTC-3′ |
| DsylCYP37 | 5′-ATGCGTTTGTACCCTTCTCG-3′ | 5′-TCGGAAATTTCTCAGGATGG-3′ |
| DsylCYP55 | 5′-CCCTGTGGGGCTGTTATAGA-3′ | 5′-CCGGAAAGAAATCTGGATGA-3′ |
| DsylGAPDH | 5′-TGCACAACTAACTGCCTTGC-3′ | 5′-TGGAAGCGGGAATTATGTTC-3′ |
| 基因名称 | ORF长度/aa | 是否有完整ORF | 等电点pI | 分子量/Da | 最佳匹配登录号 | 覆盖率/% | 相似度/% |
|---|---|---|---|---|---|---|---|
| DsylCXE1 | 72 | 否 | 10.26 | 8 104.48 | QEA03473.1 | 100 | 68.49 |
| DsylCXE2 | 132 | 否 | 9.14 | 13 850.71 | WBO25687.1 | 100 | 97.74 |
| DsylCXE3 | 69 | 否 | 8.24 | 7 756.10 | WBO25685.1 | 100 | 98.57 |
| DsylCXE4 | 569 | 是 | 5.26 | 64 239.31 | WBO25673.1 | 98 | 92.79 |
| DsylCXE5 | 146 | 否 | 4.20 | 16 401.57 | QZC92208.1 | 100 | 97.28 |
| DsylCXE6 | 543 | 是 | 6.61 | 60 808.34 | QEA03460.1 | 100 | 82.18 |
| DsylCXE7 | 135 | 否 | 9.92 | 15 497.94 | QZC92238.1 | 87 | 91.53 |
| DsylCXE8 | 469 | 是 | 4.79 | 52 641.05 | QEA03454.1 | 100 | 86.78 |
| DsylCXE9 | 269 | 是 | 5.52 | 30 268.99 | WBO25687.1 | 100 | 94.42 |
| DsylCXE10 | 65 | 否 | 11.79 | 7 408.83 | QEA03454.1 | 100 | 86.36 |
| DsylCXE11 | 340 | 是 | 6.05 | 39 054.24 | WBO25685.1 | 99 | 80.47 |
| DsylCXE12 | 202 | 是 | 4.12 | 22 542.12 | WBO25688.1 | 82 | 94.55 |
| DsylCXE13 | 209 | 是 | 10.34 | 22 976.37 | XP_053605942.1 | 96 | 57.71 |
| DsylCXE14 | 88 | 是 | 10.24 | 9 591.08 | XP_028177914.1 | 90 | 56.96 |
| DsylCXE15 | 243 | 是 | 9.30 | 26 289.89 | XP_013200959.2 | 100 | 47.98 |
| DsylCXE16 | 79 | 否 | 4.12 | 9 424.55 | XP_037294223.1 | 100 | 65.00 |
| DsylCXE17 | 476 | 是 | 5.41 | 53 153.99 | XP_053608208.1 | 100 | 72.33 |
| DsylCXE18 | 566 | 是 | 5.35 | 63 258.66 | WNK22299.1 | 99 | 71.95 |
| DsylCXE19 | 341 | 是 | 11.44 | 37 478.18 | XP_013193107.1 | 100 | 91.52 |
| DsylCXE20 | 527 | 是 | 4.84 | 59 161.58 | XP_053606293.1 | 99 | 71.51 |
| DsylCXE21 | 603 | 是 | 8.20 | 67 745.85 | XP_052759019.1 | 99 | 64.77 |
| DsylCXE22 | 550 | 是 | 7.00 | 60 391.19 | QEA03464.1 | 100 | 79.01 |
| DsylCXE23 | 546 | 是 | 6.58 | 62 718.06 | XP_013190276.2 | 100 | 76.09 |
| DsylCXE24 | 193 | 否 | 5.76 | 21 994.24 | XP_013185431.1 | 98 | 72.25 |
| DsylCXE25 | 316 | 是 | 4.47 | 35 776.48 | QEA03461.1 | 100 | 75.32 |
| DsylCXE26 | 330 | 是 | 5.96 | 38 378.05 | XP_013183096.1 | 99 | 71.65 |
| DsylCXE27 | 141 | 是 | 8.28 | 16 598.00 | XP_053622034.1 | 100 | 70.92 |
| DsylCXE28 | 532 | 是 | 6.11 | 60 081.62 | XP_053604405.1 | 100 | 77.30 |
| DsylCXE29 | 161 | 是 | 9.26 | 18 053.71 | XP_013192630.1 | 99 | 77.50 |
| DsylCXE30 | 543 | 是 | 6.31 | 61 038.43 | WBO25679.1 | 83 | 87.64 |
| DsylCXE31 | 78 | 否 | 3.98 | 9 245.44 | XDZ19301.1 | 95 | 65.33 |
| DsylCXE32 | 551 | 是 | 6.93 | 62 016.50 | QEA03463.1 | 99 | 72.86 |
| DsylCXE33 | 185 | 是 | 4.46 | 21 261.61 | XP_013196080.2 | 100 | 75.14 |
| DsylCXE34 | 340 | 是 | 8.27 | 38 859.50 | XP_053603164.1 | 100 | 74.49 |
| DsylCXE35 | 538 | 是 | 7.32 | 61 488.95 | XP_053614450.1 | 100 | 73.65 |
| DsylCXE36 | 150 | 否 | 8.99 | 16 865.08 | WBO25686.1 | 100 | 96.69 |
| DsylCXE37 | 531 | 是 | 5.14 | 59 761.73 | XP_028168560.1 | 95 | 66.67 |
| DsylCXE38 | 546 | 是 | 7.90 | 61 994.92 | QEA03466.1 | 99 | 86.64 |
| DsylCXE39 | 589 | 是 | 7.13 | 65 697.92 | WBO25685.1 | 72 | 80.84 |
表 2 赤松梢斑螟CXEs基因的筛选
Table 2 Screening of CXEs genes in Dioryctria sylvestrella
| 基因名称 | ORF长度/aa | 是否有完整ORF | 等电点pI | 分子量/Da | 最佳匹配登录号 | 覆盖率/% | 相似度/% |
|---|---|---|---|---|---|---|---|
| DsylCXE1 | 72 | 否 | 10.26 | 8 104.48 | QEA03473.1 | 100 | 68.49 |
| DsylCXE2 | 132 | 否 | 9.14 | 13 850.71 | WBO25687.1 | 100 | 97.74 |
| DsylCXE3 | 69 | 否 | 8.24 | 7 756.10 | WBO25685.1 | 100 | 98.57 |
| DsylCXE4 | 569 | 是 | 5.26 | 64 239.31 | WBO25673.1 | 98 | 92.79 |
| DsylCXE5 | 146 | 否 | 4.20 | 16 401.57 | QZC92208.1 | 100 | 97.28 |
| DsylCXE6 | 543 | 是 | 6.61 | 60 808.34 | QEA03460.1 | 100 | 82.18 |
| DsylCXE7 | 135 | 否 | 9.92 | 15 497.94 | QZC92238.1 | 87 | 91.53 |
| DsylCXE8 | 469 | 是 | 4.79 | 52 641.05 | QEA03454.1 | 100 | 86.78 |
| DsylCXE9 | 269 | 是 | 5.52 | 30 268.99 | WBO25687.1 | 100 | 94.42 |
| DsylCXE10 | 65 | 否 | 11.79 | 7 408.83 | QEA03454.1 | 100 | 86.36 |
| DsylCXE11 | 340 | 是 | 6.05 | 39 054.24 | WBO25685.1 | 99 | 80.47 |
| DsylCXE12 | 202 | 是 | 4.12 | 22 542.12 | WBO25688.1 | 82 | 94.55 |
| DsylCXE13 | 209 | 是 | 10.34 | 22 976.37 | XP_053605942.1 | 96 | 57.71 |
| DsylCXE14 | 88 | 是 | 10.24 | 9 591.08 | XP_028177914.1 | 90 | 56.96 |
| DsylCXE15 | 243 | 是 | 9.30 | 26 289.89 | XP_013200959.2 | 100 | 47.98 |
| DsylCXE16 | 79 | 否 | 4.12 | 9 424.55 | XP_037294223.1 | 100 | 65.00 |
| DsylCXE17 | 476 | 是 | 5.41 | 53 153.99 | XP_053608208.1 | 100 | 72.33 |
| DsylCXE18 | 566 | 是 | 5.35 | 63 258.66 | WNK22299.1 | 99 | 71.95 |
| DsylCXE19 | 341 | 是 | 11.44 | 37 478.18 | XP_013193107.1 | 100 | 91.52 |
| DsylCXE20 | 527 | 是 | 4.84 | 59 161.58 | XP_053606293.1 | 99 | 71.51 |
| DsylCXE21 | 603 | 是 | 8.20 | 67 745.85 | XP_052759019.1 | 99 | 64.77 |
| DsylCXE22 | 550 | 是 | 7.00 | 60 391.19 | QEA03464.1 | 100 | 79.01 |
| DsylCXE23 | 546 | 是 | 6.58 | 62 718.06 | XP_013190276.2 | 100 | 76.09 |
| DsylCXE24 | 193 | 否 | 5.76 | 21 994.24 | XP_013185431.1 | 98 | 72.25 |
| DsylCXE25 | 316 | 是 | 4.47 | 35 776.48 | QEA03461.1 | 100 | 75.32 |
| DsylCXE26 | 330 | 是 | 5.96 | 38 378.05 | XP_013183096.1 | 99 | 71.65 |
| DsylCXE27 | 141 | 是 | 8.28 | 16 598.00 | XP_053622034.1 | 100 | 70.92 |
| DsylCXE28 | 532 | 是 | 6.11 | 60 081.62 | XP_053604405.1 | 100 | 77.30 |
| DsylCXE29 | 161 | 是 | 9.26 | 18 053.71 | XP_013192630.1 | 99 | 77.50 |
| DsylCXE30 | 543 | 是 | 6.31 | 61 038.43 | WBO25679.1 | 83 | 87.64 |
| DsylCXE31 | 78 | 否 | 3.98 | 9 245.44 | XDZ19301.1 | 95 | 65.33 |
| DsylCXE32 | 551 | 是 | 6.93 | 62 016.50 | QEA03463.1 | 99 | 72.86 |
| DsylCXE33 | 185 | 是 | 4.46 | 21 261.61 | XP_013196080.2 | 100 | 75.14 |
| DsylCXE34 | 340 | 是 | 8.27 | 38 859.50 | XP_053603164.1 | 100 | 74.49 |
| DsylCXE35 | 538 | 是 | 7.32 | 61 488.95 | XP_053614450.1 | 100 | 73.65 |
| DsylCXE36 | 150 | 否 | 8.99 | 16 865.08 | WBO25686.1 | 100 | 96.69 |
| DsylCXE37 | 531 | 是 | 5.14 | 59 761.73 | XP_028168560.1 | 95 | 66.67 |
| DsylCXE38 | 546 | 是 | 7.90 | 61 994.92 | QEA03466.1 | 99 | 86.64 |
| DsylCXE39 | 589 | 是 | 7.13 | 65 697.92 | WBO25685.1 | 72 | 80.84 |
| 基因名称 | ORF长度/aa | 是否有完整ORF | 等电点pI | 分子量/Da | 最佳匹配登录号 | 覆盖率/% | 相似度/% |
|---|---|---|---|---|---|---|---|
| DsylAOX1 | 133 | 否 | 4.94 | 15 161.26 | QZC92247.1 | 100 | 97.01 |
| DsylAOX2 | 1 028 | 是 | 5.33 | 115 397.04 | NP_001299599.1 | 100 | 79.57 |
| DsylAOX3 | 134 | 否 | 8.42 | 14 787.10 | NP_001299599.1 | 96 | 91.47 |
| DsylAOX4 | 1 280 | 是 | 6.39 | 142 514.41 | BAR64767.1 | 100 | 72.76 |
| DsylAOX5 | 589 | 是 | 9.97 | 66 171.90 | QPF77600.1 | 68 | 63.75 |
| DsylAOX6 | 372 | 否 | 5.33 | 42 054.92 | XP_053608971.1 | 100 | 80.97 |
表 3 赤松梢斑螟AOXs基因的筛选
Table 3 Screening of AOXs genes in Dioryctria sylvestrella
| 基因名称 | ORF长度/aa | 是否有完整ORF | 等电点pI | 分子量/Da | 最佳匹配登录号 | 覆盖率/% | 相似度/% |
|---|---|---|---|---|---|---|---|
| DsylAOX1 | 133 | 否 | 4.94 | 15 161.26 | QZC92247.1 | 100 | 97.01 |
| DsylAOX2 | 1 028 | 是 | 5.33 | 115 397.04 | NP_001299599.1 | 100 | 79.57 |
| DsylAOX3 | 134 | 否 | 8.42 | 14 787.10 | NP_001299599.1 | 96 | 91.47 |
| DsylAOX4 | 1 280 | 是 | 6.39 | 142 514.41 | BAR64767.1 | 100 | 72.76 |
| DsylAOX5 | 589 | 是 | 9.97 | 66 171.90 | QPF77600.1 | 68 | 63.75 |
| DsylAOX6 | 372 | 否 | 5.33 | 42 054.92 | XP_053608971.1 | 100 | 80.97 |
| 基因名称 | ORF长度/aa | 是否有完整ORF | 等电点pI | 分子量/Da | 最佳匹配登录号 | 覆盖率/% | 相似度/% |
|---|---|---|---|---|---|---|---|
| DsylGST1 | 97 | 否 | 4.49 | 11 307.85 | WBO25703.1 | 100 | 92.86 |
| DsylGST2 | 68 | 否 | 4.78 | 7 828.27 | WBO25700.1 | 100 | 98.55 |
| DsylGST3 | 65 | 否 | 5.72 | 7 661.77 | WBO25700.1 | 100 | 96.97 |
| DsylGST4 | 57 | 否 | 7.51 | 6 505.81 | WBO25700.1 | 100 | 100.00 |
| DsylGST5 | 101 | 是 | 9.16 | 11 257.77 | XP_053614950.1 | 95 | 83.33 |
| DsylGST6 | 244 | 是 | 9.12 | 28 188.68 | XP_053614946.1 | 100 | 73.77 |
| DsylGST7 | 68 | 否 | 6.95 | 7 760.02 | WBO25698.1 | 100 | 100.00 |
| DsylGST8 | 235 | 是 | 6.80 | 27 007.98 | XP_060802045.1 | 99 | 87.98 |
| DsylGST9 | 68 | 否 | 9.98 | 7 803.42 | NP_001299595.1 | 100 | 60.87 |
| DsylGST10 | 241 | 是 | 5.76 | 27 083.19 | WBO25700.1 | 98 | 96.19 |
| DsylGST11 | 289 | 是 | 7.23 | 33 226.65 | WBO25693.1 | 100 | 89.27 |
| DsylGST12 | 221 | 是 | 6.53 | 24 972.69 | XP_053614426.1 | 97 | 85.12 |
| DsylGST13 | 219 | 是 | 5.86 | 24 718.29 | WBO25696.1 | 87 | 95.29 |
| DsylGST14 | 246 | 是 | 5.32 | 27 579.84 | NP_001299595.1 | 100 | 89.02 |
| DsylGST15 | 216 | 是 | 4.93 | 24 438.09 | UQS35834.1 | 100 | 78.60 |
| DsylGST16 | 255 | 是 | 6.94 | 28 903.33 | WBO25695.1 | 100 | 98.82 |
| DsylGST17 | 225 | 是 | 9.18 | 25 376.48 | WBO25699.1 | 91 | 93.66 |
| DsylGST18 | 151 | 是 | 9.96 | 16 786.68 | XP_053625608.1 | 100 | 86.93 |
| DsylGST19 | 70 | 否 | 9.16 | 7 834.26 | WBO25700.1 | 100 | 97.18 |
| DsylGST20 | 204 | 是 | 6.30 | 23 546.20 | WBO25701.1 | 100 | 94.61 |
| DsylGST21 | 228 | 是 | 9.13 | 26 640.70 | XP_013194289.1 | 100 | 82.46 |
| DsylGST22 | 151 | 是 | 10.10 | 16 985.24 | WBO25691.1 | 77 | 94.83 |
| DsylGST23 | 214 | 是 | 7.69 | 24 064.50 | WBO25698.1 | 100 | 97.20 |
| DsylGST24 | 69 | 否 | 6.51 | 8 063.24 | WBO25700.1 | 100 | 94.29 |
| DsylGST25 | 74 | 是 | 5.03 | 8 782.15 | WBO25693.1 | 100 | 81.08 |
| DsylGST26 | 148 | 是 | 10.16 | 16 566.47 | WBO25697.1 | 100 | 98.65 |
| DsylGST27 | 299 | 是 | 7.04 | 26 110.28 | XP_053611606.1 | 100 | 76.86 |
| DsylGST28 | 40 | 否 | 10.96 | 4 455.30 | NP_001299595.1 | 100 | 80.49 |
| DsylGST29 | 113 | 否 | 5.07 | 12 937.99 | WBO25700.1 | 94 | 72.48 |
| DsylGST30 | 204 | 是 | 4.93 | 23 676.35 | WBO25703.1 | 78 | 100.00 |
| DsylGST31 | 248 | 是 | 8.75 | 28 661.32 | WBO25692.1 | 86 | 99.53 |
| DsylGST32 | 78 | 否 | 9.39 | 8 883.29 | QCX41803.1 | 99 | 74.36 |
表 4 赤松梢斑螟GSTs基因的筛选
Table 4 Screening of GSTs genes in Dioryctria sylvestrella
| 基因名称 | ORF长度/aa | 是否有完整ORF | 等电点pI | 分子量/Da | 最佳匹配登录号 | 覆盖率/% | 相似度/% |
|---|---|---|---|---|---|---|---|
| DsylGST1 | 97 | 否 | 4.49 | 11 307.85 | WBO25703.1 | 100 | 92.86 |
| DsylGST2 | 68 | 否 | 4.78 | 7 828.27 | WBO25700.1 | 100 | 98.55 |
| DsylGST3 | 65 | 否 | 5.72 | 7 661.77 | WBO25700.1 | 100 | 96.97 |
| DsylGST4 | 57 | 否 | 7.51 | 6 505.81 | WBO25700.1 | 100 | 100.00 |
| DsylGST5 | 101 | 是 | 9.16 | 11 257.77 | XP_053614950.1 | 95 | 83.33 |
| DsylGST6 | 244 | 是 | 9.12 | 28 188.68 | XP_053614946.1 | 100 | 73.77 |
| DsylGST7 | 68 | 否 | 6.95 | 7 760.02 | WBO25698.1 | 100 | 100.00 |
| DsylGST8 | 235 | 是 | 6.80 | 27 007.98 | XP_060802045.1 | 99 | 87.98 |
| DsylGST9 | 68 | 否 | 9.98 | 7 803.42 | NP_001299595.1 | 100 | 60.87 |
| DsylGST10 | 241 | 是 | 5.76 | 27 083.19 | WBO25700.1 | 98 | 96.19 |
| DsylGST11 | 289 | 是 | 7.23 | 33 226.65 | WBO25693.1 | 100 | 89.27 |
| DsylGST12 | 221 | 是 | 6.53 | 24 972.69 | XP_053614426.1 | 97 | 85.12 |
| DsylGST13 | 219 | 是 | 5.86 | 24 718.29 | WBO25696.1 | 87 | 95.29 |
| DsylGST14 | 246 | 是 | 5.32 | 27 579.84 | NP_001299595.1 | 100 | 89.02 |
| DsylGST15 | 216 | 是 | 4.93 | 24 438.09 | UQS35834.1 | 100 | 78.60 |
| DsylGST16 | 255 | 是 | 6.94 | 28 903.33 | WBO25695.1 | 100 | 98.82 |
| DsylGST17 | 225 | 是 | 9.18 | 25 376.48 | WBO25699.1 | 91 | 93.66 |
| DsylGST18 | 151 | 是 | 9.96 | 16 786.68 | XP_053625608.1 | 100 | 86.93 |
| DsylGST19 | 70 | 否 | 9.16 | 7 834.26 | WBO25700.1 | 100 | 97.18 |
| DsylGST20 | 204 | 是 | 6.30 | 23 546.20 | WBO25701.1 | 100 | 94.61 |
| DsylGST21 | 228 | 是 | 9.13 | 26 640.70 | XP_013194289.1 | 100 | 82.46 |
| DsylGST22 | 151 | 是 | 10.10 | 16 985.24 | WBO25691.1 | 77 | 94.83 |
| DsylGST23 | 214 | 是 | 7.69 | 24 064.50 | WBO25698.1 | 100 | 97.20 |
| DsylGST24 | 69 | 否 | 6.51 | 8 063.24 | WBO25700.1 | 100 | 94.29 |
| DsylGST25 | 74 | 是 | 5.03 | 8 782.15 | WBO25693.1 | 100 | 81.08 |
| DsylGST26 | 148 | 是 | 10.16 | 16 566.47 | WBO25697.1 | 100 | 98.65 |
| DsylGST27 | 299 | 是 | 7.04 | 26 110.28 | XP_053611606.1 | 100 | 76.86 |
| DsylGST28 | 40 | 否 | 10.96 | 4 455.30 | NP_001299595.1 | 100 | 80.49 |
| DsylGST29 | 113 | 否 | 5.07 | 12 937.99 | WBO25700.1 | 94 | 72.48 |
| DsylGST30 | 204 | 是 | 4.93 | 23 676.35 | WBO25703.1 | 78 | 100.00 |
| DsylGST31 | 248 | 是 | 8.75 | 28 661.32 | WBO25692.1 | 86 | 99.53 |
| DsylGST32 | 78 | 否 | 9.39 | 8 883.29 | QCX41803.1 | 99 | 74.36 |
| 基因名称 | ORF长度/aa | 是否有完整ORF | 等电点pI | 分子量/Da | 最佳匹配登录号 | 覆盖率/% | 相似度/% |
|---|---|---|---|---|---|---|---|
| DsylCYP1 | 25 | 否 | 4.37 | 2 811.33 | XP_060801165.1 | 100 | 69.23 |
| DsylCYP2 | 260 | 否 | 5.26 | 30 196.61 | QZP43566.1 | 100 | 77.01 |
| DsylCYP3 | 203 | 否 | 7.40 | 23 604.40 | XP_053613700.1 | 100 | 87.75 |
| DsylCYP4 | 232 | 否 | 6.79 | 26 583.99 | XP_064292090.1 | 100 | 80.60 |
| DsylCYP5 | 79 | 否 | 3.87 | 9 071.09 | ULR85505.1 | 100 | 86.25 |
| DsylCYP6 | 61 | 否 | 10.48 | 7 354.68 | XP_060805041.1 | 100 | 70.97 |
| DsylCYP7 | 64 | 否 | 4.86 | 7 457.61 | XP_013183008.2 | 100 | 84.62 |
| DsylCYP8 | 68 | 否 | 4.36 | 7 856.90 | WBO25717.1 | 100 | 86.96 |
| DsylCYP9 | 120 | 否 | 4.20 | 14 186.11 | WBO25708.1 | 98 | 94.12 |
| DsylCYP10 | 109 | 否 | 6.09 | 12 554.26 | GLH02031.1 | 100 | 100.00 |
| DsylCYP11 | 43 | 否 | 4.24 | 4 887.45 | QPF77601.1 | 100 | 95.45 |
| DsylCYP12 | 90 | 否 | 7.96 | 10 338.05 | XP_060800329.1 | 100 | 80.22 |
| DsylCYP13 | 63 | 否 | 5.70 | 7 164.29 | QZP43568.1 | 98 | 90.48 |
| DsylCYP14 | 499 | 是 | 7.91 | 57 745.26 | WBO25716.1 | 100 | 90.36 |
| DsylCYP15 | 50 | 否 | 10.60 | 6 359.78 | XP_013196728.2 | 98 | 82.00 |
| DsylCYP16 | 143 | 是 | 10.51 | 16 007.83 | QZP43575.1 | 100 | 88.11 |
| DsylCYP17 | 383 | 否 | 6.34 | 45 061.09 | XP_013183823.2 | 96 | 72.90 |
| DsylCYP18 | 593 | 是 | 6.51 | 61 766.51 | QZP43558.1 | 100 | 86.27 |
| DsylCYP19 | 511 | 是 | 8.38 | 58 470.84 | XP_013186352.1 | 100 | 74.46 |
| DsylCYP20 | 531 | 是 | 8.06 | 60 900.52 | QZP43566.1 | 100 | 72.69 |
| DsylCYP21 | 481 | 是 | 6.34 | 56 412.28 | NP_001351990.1 | 99 | 78.48 |
| DsylCYP22 | 531 | 是 | 9.20 | 61 763.66 | XP_013199514.1 | 100 | 86.23 |
| DsylCYP23 | 505 | 是 | 9.03 | 57 711.08 | XP_060804943.1 | 98 | 740.4 |
| DsylCYP24 | 513 | 是 | 8.44 | 58 529.74 | XP_013184491.1 | 100 | 82.26 |
| DsylCYP25 | 502 | 是 | 9.05 | 58 062.36 | XP_013183008.2 | 99 | 74.25 |
| DsylCYP26 | 531 | 是 | 9.55 | 60 886.97 | XP_053625564.1 | 98 | 64.15 |
| DsylCYP27 | 197 | 否 | 9.51 | 23 097.04 | WBO25718.1 | 100 | 94.44 |
| DsylCYP28 | 513 | 是 | 9.35 | 58 743.97 | QZP43553.1 | 100 | 75.88 |
| DsylCYP29 | 498 | 是 | 9.06 | 57 031.47 | XP_013192856.2 | 100 | 80.92 |
| DsylCYP30 | 517 | 是 | 8.92 | 59 152.29 | QZP43555.1 | 98 | 79.13 |
| DsylCYP31 | 515 | 是 | 9.66 | 59 588.69 | XP_013184001.2 | 99 | 81.45 |
| DsylCYP32 | 502 | 是 | 9.25 | 58 599.23 | XP_013183823.2 | 97 | 70.70 |
| DsylCYP33 | 515 | 是 | 9.00 | 58 784.25 | WBO25715.1 | 99 | 96.88 |
| DsylCYP34 | 512 | 是 | 8.82 | 59 185.80 | WBO25709.1 | 98 | 96.83 |
| DsylCYP35 | 412 | 否 | 9.55 | 47 467.23 | XP_013188296.2 | 99 | 54.99 |
| DsylCYP36 | 510 | 是 | 9.25 | 57 968.50 | XP_060801165.1 | 97 | 62.87 |
| DsylCYP37 | 559 | 是 | 9.08 | 63 964.16 | WBO25721.1 | 100 | 97.13 |
| DsylCYP38 | 524 | 是 | 8.86 | 61 057.87 | XP_060802800.1 | 99 | 78.08 |
| DsylCYP39 | 684 | 是 | 5.22 | 76 991.49 | WBO25714.1 | 100 | 96.35 |
| DsylCYP40 | 439 | 是 | 9.74 | 50 541.80 | XP_013188296.2 | 100 | 55.91 |
| DsylCYP41 | 522 | 是 | 8.53 | 61 340.15 | XP_060806632.1 | 99 | 78.61 |
| DsylCYP42 | 493 | 是 | 8.67 | 57 387.32 | XP_053601586.1 | 97 | 77.41 |
| DsylCYP43 | 481 | 是 | 6.72 | 53 682.25 | XP_013183200.1 | 100 | 86.07 |
| DsylCYP44 | 540 | 是 | 7.52 | 61 379.28 | QZP43565.1 | 100 | 95.37 |
| DsylCYP45 | 87 | 是 | 10.19 | 10 410.61 | XP_013185828.2 | 36 | 100.00 |
| DsylCYP46 | 258 | 是 | 8.77 | 29 618.27 | XP_059050485.1 | 100 | 54.82 |
| DsylCYP47 | 515 | 是 | 8.71 | 59 383.15 | XP_060803177.1 | 100 | 86.27 |
| DsylCYP48 | 510 | 是 | 8.28 | 59 295.82 | QZP43571.1 | 100 | 79.80 |
| DsylCYP49 | 45 | 否 | 8.93 | 5 504.43 | XP_013185828.2 | 100 | 93.48 |
| DsylCYP50 | 103 | 是 | 11.06 | 12 188.29 | QBA57446.1 | 96 | 74.75 |
| DsylCYP51 | 235 | 是 | 7.86 | 27 033.33 | QZP43554.1 | 95 | 72.65 |
| DsylCYP52 | 509 | 是 | 9.71 | 57 937.54 | XP_013192036.1 | 100 | 77.76 |
| DsylCYP53 | 522 | 是 | 8.30 | 61 249.98 | XP_060806632.1 | 99 | 78.61 |
| DsylCYP54 | 48 | 是 | 9.97 | 5 665.76 | XP_060804943.1 | 94 | 66.67 |
| DsylCYP55 | 521 | 是 | 7.24 | 60 774.40 | XP_053617327.1 | 100 | 76.15 |
| DsylCYP56 | 169 | 是 | 10.10 | 19 784.42 | XP_013188296.2 | 41 | 69.44 |
| DsylCYP57 | 66 | 是 | 10.82 | 7 665.25 | XP_013199514.1 | 98 | 92.31 |
| DsylCYP58 | 166 | 是 | 6.11 | 18 856.61 | XP_013189127.2 | 95 | 89.17 |
| DsylCYP59 | 244 | 否 | 7.03 | 28 080.67 | QBA57484.1 | 100 | 96.72 |
| DsylCYP60 | 507 | 是 | 8.58 | 58 260.37 | XP_053613686.1 | 100 | 69.57 |
表 5 赤松梢斑螟CYPs基因的筛选
Table 5 Screening of CYPs genes in Dioryctria sylvestrella
| 基因名称 | ORF长度/aa | 是否有完整ORF | 等电点pI | 分子量/Da | 最佳匹配登录号 | 覆盖率/% | 相似度/% |
|---|---|---|---|---|---|---|---|
| DsylCYP1 | 25 | 否 | 4.37 | 2 811.33 | XP_060801165.1 | 100 | 69.23 |
| DsylCYP2 | 260 | 否 | 5.26 | 30 196.61 | QZP43566.1 | 100 | 77.01 |
| DsylCYP3 | 203 | 否 | 7.40 | 23 604.40 | XP_053613700.1 | 100 | 87.75 |
| DsylCYP4 | 232 | 否 | 6.79 | 26 583.99 | XP_064292090.1 | 100 | 80.60 |
| DsylCYP5 | 79 | 否 | 3.87 | 9 071.09 | ULR85505.1 | 100 | 86.25 |
| DsylCYP6 | 61 | 否 | 10.48 | 7 354.68 | XP_060805041.1 | 100 | 70.97 |
| DsylCYP7 | 64 | 否 | 4.86 | 7 457.61 | XP_013183008.2 | 100 | 84.62 |
| DsylCYP8 | 68 | 否 | 4.36 | 7 856.90 | WBO25717.1 | 100 | 86.96 |
| DsylCYP9 | 120 | 否 | 4.20 | 14 186.11 | WBO25708.1 | 98 | 94.12 |
| DsylCYP10 | 109 | 否 | 6.09 | 12 554.26 | GLH02031.1 | 100 | 100.00 |
| DsylCYP11 | 43 | 否 | 4.24 | 4 887.45 | QPF77601.1 | 100 | 95.45 |
| DsylCYP12 | 90 | 否 | 7.96 | 10 338.05 | XP_060800329.1 | 100 | 80.22 |
| DsylCYP13 | 63 | 否 | 5.70 | 7 164.29 | QZP43568.1 | 98 | 90.48 |
| DsylCYP14 | 499 | 是 | 7.91 | 57 745.26 | WBO25716.1 | 100 | 90.36 |
| DsylCYP15 | 50 | 否 | 10.60 | 6 359.78 | XP_013196728.2 | 98 | 82.00 |
| DsylCYP16 | 143 | 是 | 10.51 | 16 007.83 | QZP43575.1 | 100 | 88.11 |
| DsylCYP17 | 383 | 否 | 6.34 | 45 061.09 | XP_013183823.2 | 96 | 72.90 |
| DsylCYP18 | 593 | 是 | 6.51 | 61 766.51 | QZP43558.1 | 100 | 86.27 |
| DsylCYP19 | 511 | 是 | 8.38 | 58 470.84 | XP_013186352.1 | 100 | 74.46 |
| DsylCYP20 | 531 | 是 | 8.06 | 60 900.52 | QZP43566.1 | 100 | 72.69 |
| DsylCYP21 | 481 | 是 | 6.34 | 56 412.28 | NP_001351990.1 | 99 | 78.48 |
| DsylCYP22 | 531 | 是 | 9.20 | 61 763.66 | XP_013199514.1 | 100 | 86.23 |
| DsylCYP23 | 505 | 是 | 9.03 | 57 711.08 | XP_060804943.1 | 98 | 740.4 |
| DsylCYP24 | 513 | 是 | 8.44 | 58 529.74 | XP_013184491.1 | 100 | 82.26 |
| DsylCYP25 | 502 | 是 | 9.05 | 58 062.36 | XP_013183008.2 | 99 | 74.25 |
| DsylCYP26 | 531 | 是 | 9.55 | 60 886.97 | XP_053625564.1 | 98 | 64.15 |
| DsylCYP27 | 197 | 否 | 9.51 | 23 097.04 | WBO25718.1 | 100 | 94.44 |
| DsylCYP28 | 513 | 是 | 9.35 | 58 743.97 | QZP43553.1 | 100 | 75.88 |
| DsylCYP29 | 498 | 是 | 9.06 | 57 031.47 | XP_013192856.2 | 100 | 80.92 |
| DsylCYP30 | 517 | 是 | 8.92 | 59 152.29 | QZP43555.1 | 98 | 79.13 |
| DsylCYP31 | 515 | 是 | 9.66 | 59 588.69 | XP_013184001.2 | 99 | 81.45 |
| DsylCYP32 | 502 | 是 | 9.25 | 58 599.23 | XP_013183823.2 | 97 | 70.70 |
| DsylCYP33 | 515 | 是 | 9.00 | 58 784.25 | WBO25715.1 | 99 | 96.88 |
| DsylCYP34 | 512 | 是 | 8.82 | 59 185.80 | WBO25709.1 | 98 | 96.83 |
| DsylCYP35 | 412 | 否 | 9.55 | 47 467.23 | XP_013188296.2 | 99 | 54.99 |
| DsylCYP36 | 510 | 是 | 9.25 | 57 968.50 | XP_060801165.1 | 97 | 62.87 |
| DsylCYP37 | 559 | 是 | 9.08 | 63 964.16 | WBO25721.1 | 100 | 97.13 |
| DsylCYP38 | 524 | 是 | 8.86 | 61 057.87 | XP_060802800.1 | 99 | 78.08 |
| DsylCYP39 | 684 | 是 | 5.22 | 76 991.49 | WBO25714.1 | 100 | 96.35 |
| DsylCYP40 | 439 | 是 | 9.74 | 50 541.80 | XP_013188296.2 | 100 | 55.91 |
| DsylCYP41 | 522 | 是 | 8.53 | 61 340.15 | XP_060806632.1 | 99 | 78.61 |
| DsylCYP42 | 493 | 是 | 8.67 | 57 387.32 | XP_053601586.1 | 97 | 77.41 |
| DsylCYP43 | 481 | 是 | 6.72 | 53 682.25 | XP_013183200.1 | 100 | 86.07 |
| DsylCYP44 | 540 | 是 | 7.52 | 61 379.28 | QZP43565.1 | 100 | 95.37 |
| DsylCYP45 | 87 | 是 | 10.19 | 10 410.61 | XP_013185828.2 | 36 | 100.00 |
| DsylCYP46 | 258 | 是 | 8.77 | 29 618.27 | XP_059050485.1 | 100 | 54.82 |
| DsylCYP47 | 515 | 是 | 8.71 | 59 383.15 | XP_060803177.1 | 100 | 86.27 |
| DsylCYP48 | 510 | 是 | 8.28 | 59 295.82 | QZP43571.1 | 100 | 79.80 |
| DsylCYP49 | 45 | 否 | 8.93 | 5 504.43 | XP_013185828.2 | 100 | 93.48 |
| DsylCYP50 | 103 | 是 | 11.06 | 12 188.29 | QBA57446.1 | 96 | 74.75 |
| DsylCYP51 | 235 | 是 | 7.86 | 27 033.33 | QZP43554.1 | 95 | 72.65 |
| DsylCYP52 | 509 | 是 | 9.71 | 57 937.54 | XP_013192036.1 | 100 | 77.76 |
| DsylCYP53 | 522 | 是 | 8.30 | 61 249.98 | XP_060806632.1 | 99 | 78.61 |
| DsylCYP54 | 48 | 是 | 9.97 | 5 665.76 | XP_060804943.1 | 94 | 66.67 |
| DsylCYP55 | 521 | 是 | 7.24 | 60 774.40 | XP_053617327.1 | 100 | 76.15 |
| DsylCYP56 | 169 | 是 | 10.10 | 19 784.42 | XP_013188296.2 | 41 | 69.44 |
| DsylCYP57 | 66 | 是 | 10.82 | 7 665.25 | XP_013199514.1 | 98 | 92.31 |
| DsylCYP58 | 166 | 是 | 6.11 | 18 856.61 | XP_013189127.2 | 95 | 89.17 |
| DsylCYP59 | 244 | 否 | 7.03 | 28 080.67 | QBA57484.1 | 100 | 96.72 |
| DsylCYP60 | 507 | 是 | 8.58 | 58 260.37 | XP_053613686.1 | 100 | 69.57 |
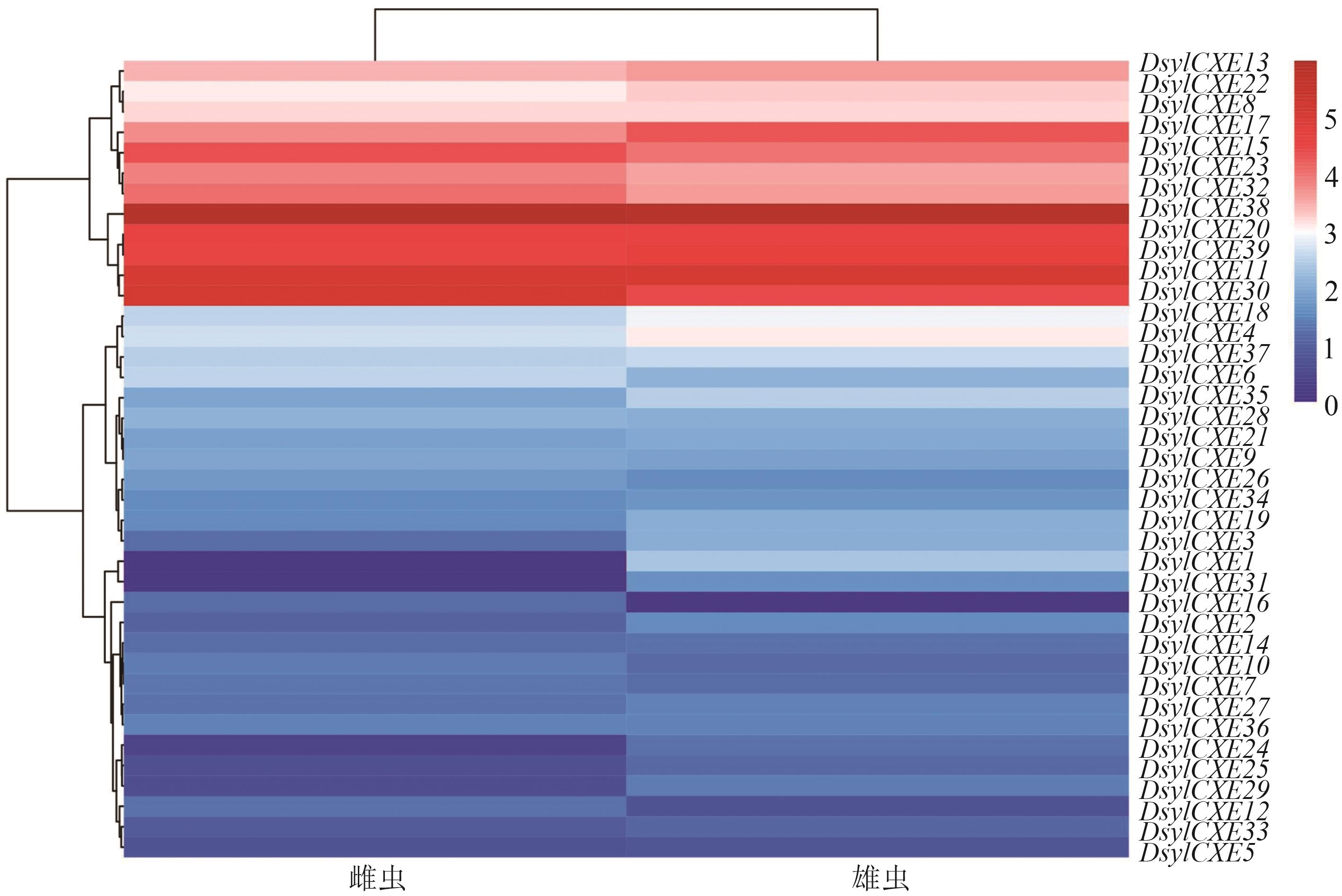
图 5 DsylCXEs在赤松梢斑螟雌雄虫触角转录组中的表达水平注:不同颜色代表基因的相对丰度值大小,颜色越红代表表达量越高。
Fig. 5 Transcript abundances of DsylCXEs in female and male antennae of D. sylvestrella
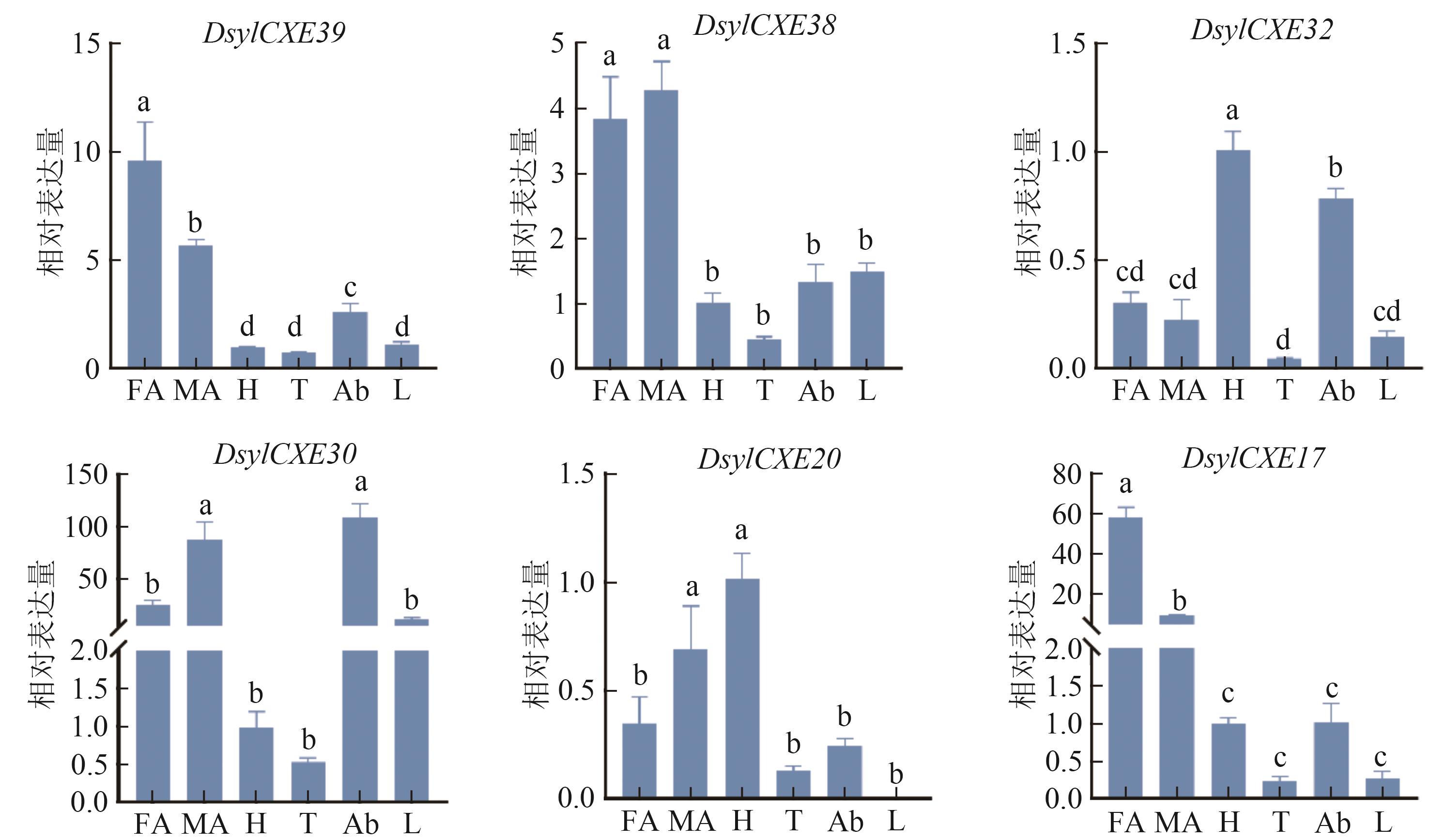
图 6 赤松梢斑螟CXEs基因在成虫不同组织中的相对表达水平注:FA—雌虫触角;MA—雄虫触角;H—头;T—胸;Ab—腹;L—足。同一图内不同小写字母表示在P0.05水平有统计学差异。
Fig. 6 The relative expression levels of CXEs genes in different adult tissues of D. sylvestrella
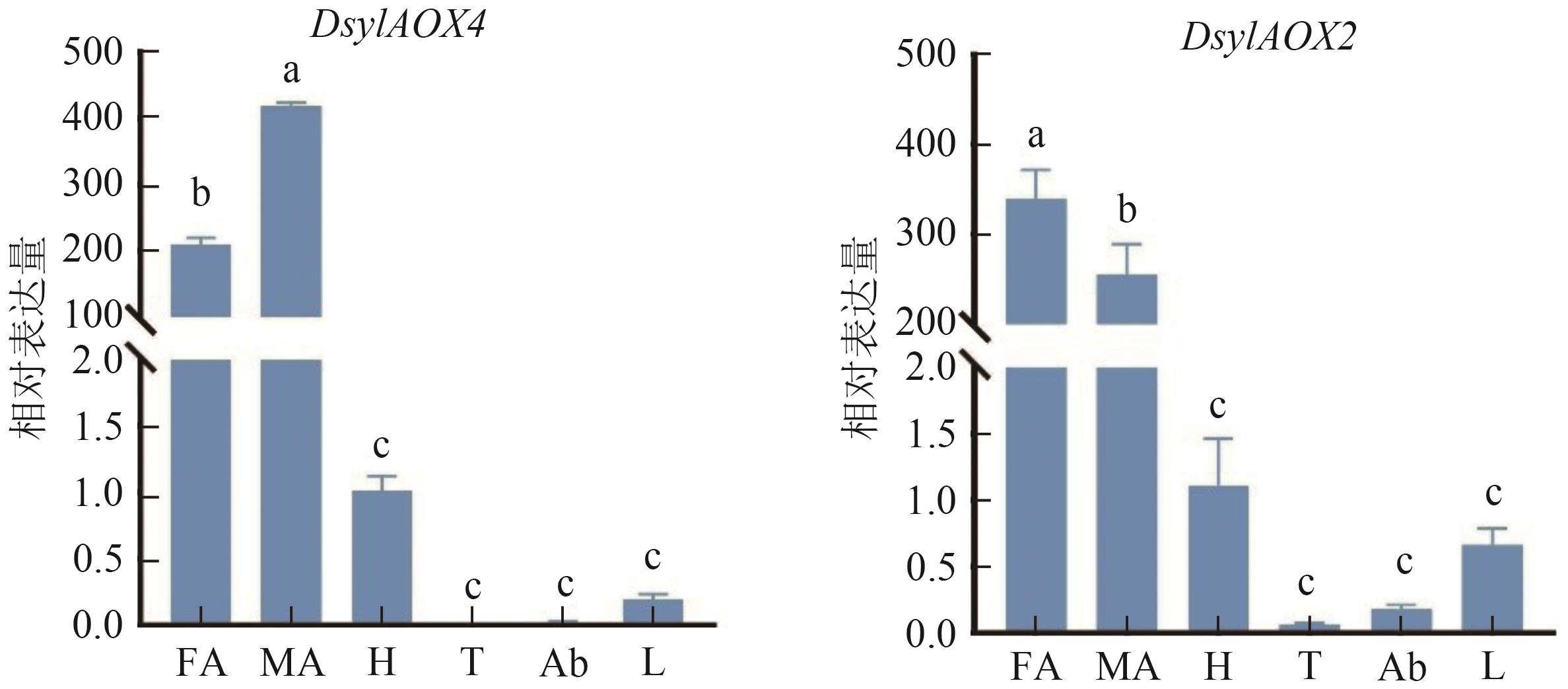
图 8 赤松梢斑螟AOXs基因在成虫不同组织中的相对表达水平注:FA—雌虫触角;MA—雄虫触角;H—头;T—胸;Ab—腹;L—足。同一图内不同小写字母表示在P0.05水平有统计学差异。
Fig. 8 The relative expression levels of AOXs genes in different adult tissues of D. sylvestrella
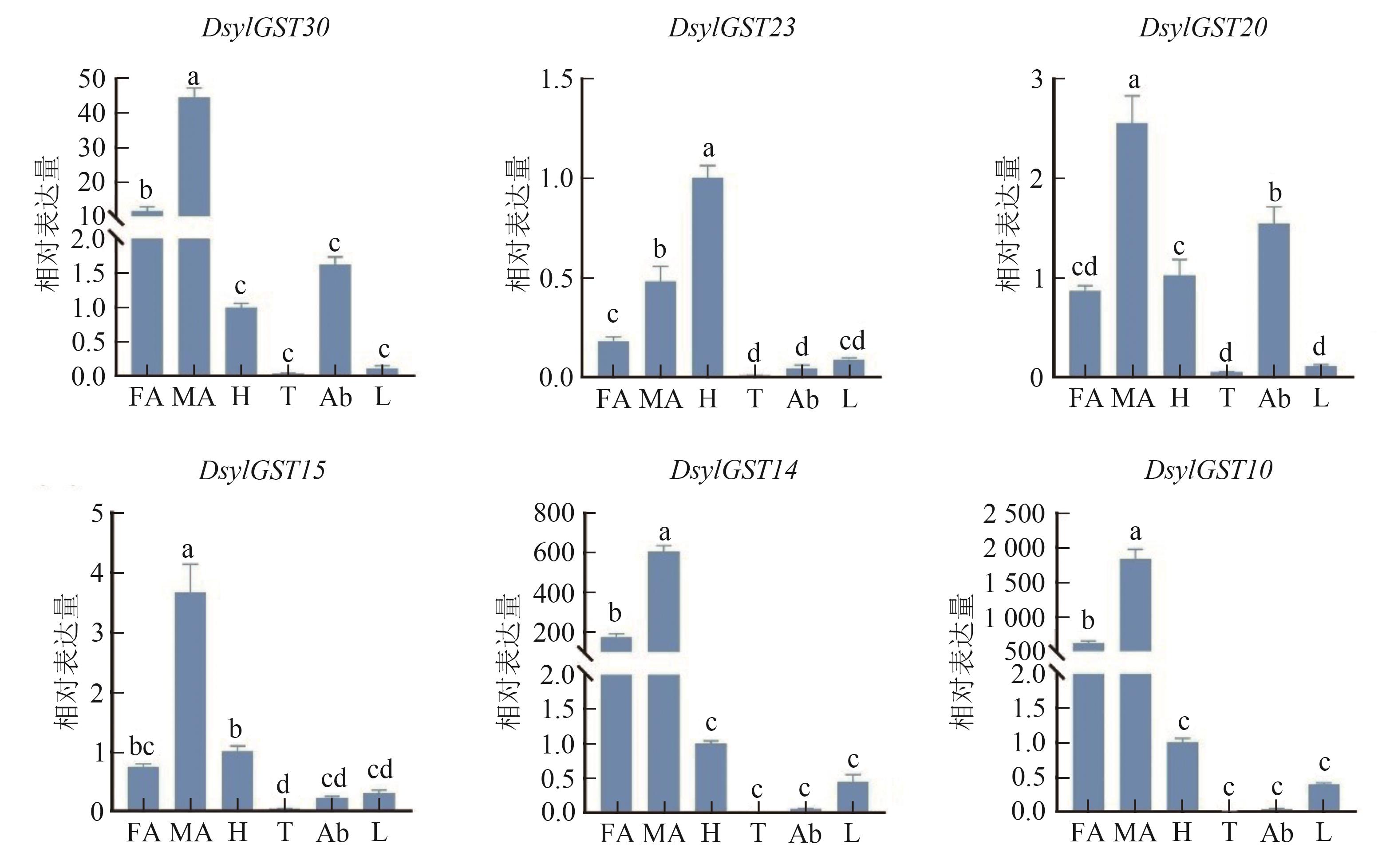
图 10 赤松梢斑螟GSTs基因在成虫不同组织中的相对表达水平注:FA—雌虫触角;MA—雄虫触角;H—头;T—胸;Ab—腹;L—足。同一图内不同小写字母表示在P0.05水平有统计学差异。
Fig. 10 The relative expression levels of GSTs genes in different adult tissues of D. sylvestrella
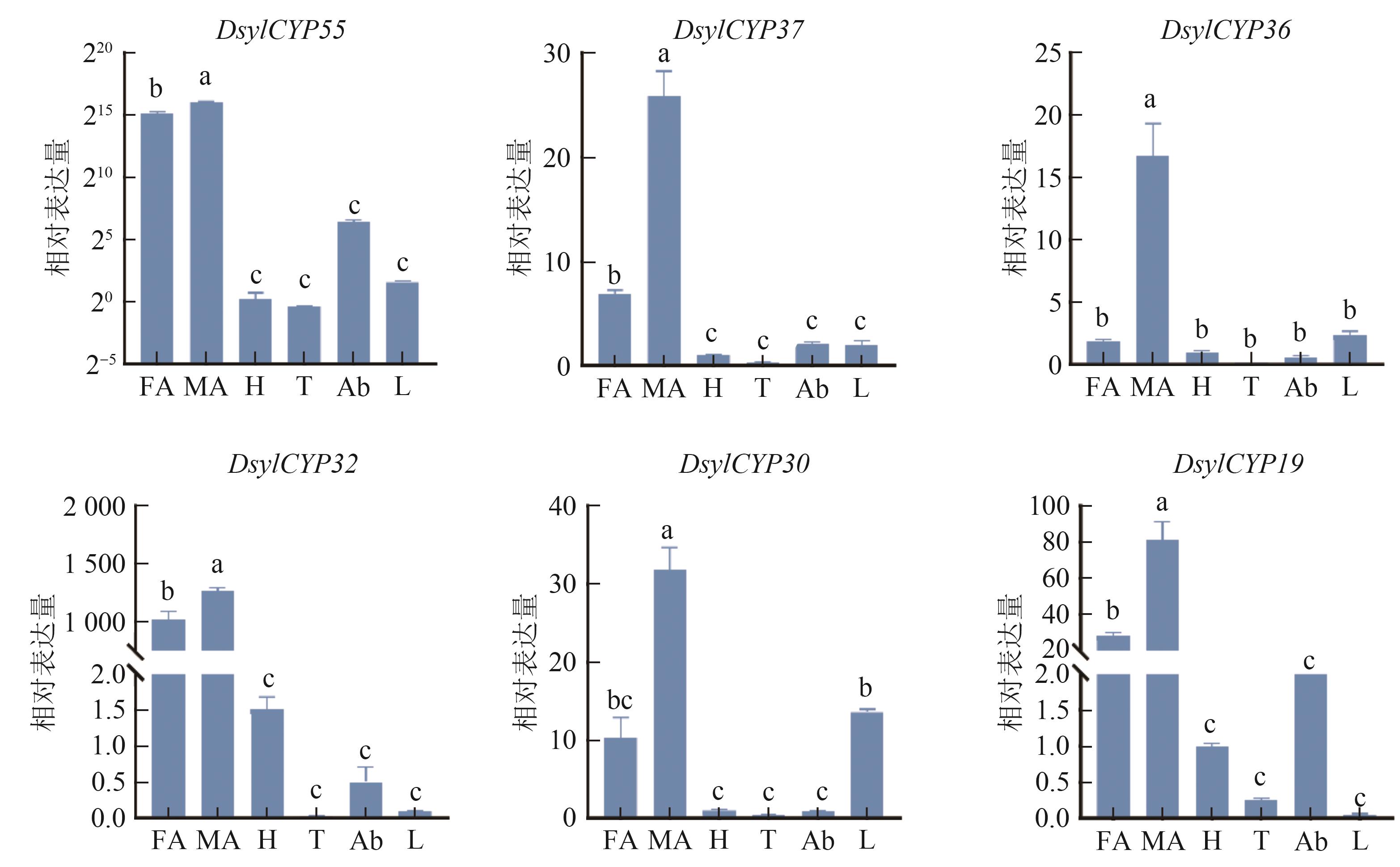
图 12 赤松梢斑螟CYPs基因在成虫不同组织中的相对表达水平注:FA—雌虫触角;MA—雄虫触角;H—头;T—胸;Ab—腹;L—足。同一图内不同小写字母表示在P0.05水平有统计学差异。
Fig. 12 The relative expression levels of CYPs genes in different adult tissues of D. sylvestrella
| [1] | 卢孟.蛀害红松和樟子松的害虫种类鉴定及遗传多样性研究[D].哈尔滨:东北林业大学, 2020. |
| [2] | 董金宝.赤松梢斑螟的发生危害及防控技术[J].黑龙江生态工程职业学院学报,2010(2):34-35. |
| DONG J B. Occurrence, harm and prevention and control techniques of Chilostachys koraiensis [J]. J. Heilongjiang Voc. Instit. Ecolog. Engin., 2010(2): 34-35. | |
| [3] | 杨香茹.红松经济林培育中出现的常见问题及发展途径[J].现代园艺,2024(10):188-189. |
| YANG X R. Common problems and development approaches in cultivation of Korean pine economic forest[J]. Contemp. Hortic., 2024(10): 188-189. | |
| [4] | 周彪,邓勋,孙家宝,等.红松种实害虫生物学特性的研究[J].防护林科技,2006(6):38-40. |
| ZHOU B, DENG X, SUN J B, et al.. Study on biological characteristics of seed pests of Korean pine[J]. Protect. Forest Sci. Technol., 2006(6): 38-40. | |
| [5] | 郭鸿儒.球孢白僵菌菌株BbSBM03对赤松梢斑螟幼虫杀虫活性及机制研究[D].哈尔滨:东北林业大学, 2024. |
| [6] | 宋玉双,李娟,张天栋,等.我国梢斑螟属害虫经济重要性分析[J].中国森林病虫,2021,40(1):1-5. |
| SONG Y S, LI J, ZHANG T D, et al.. Analysis of the economic importance of Dioryctria pests in China[J]. For. Pest Dis., 2021, 40(1): 1-5. | |
| [7] | PELOSI P. Perireceptor events in olfaction[J]. J. Neurobiol., 1996, 30(1): 3-19. |
| [8] | VOGT G R. Insect pheromone biochemistry and molecular biology: the biosynthesis and detection of pheromones and plant volatiles[M]. Elsevier: Academic Press, 2003, 391-445. |
| [9] | VOGT R G, RIDDIFORD L M. Pheromone binding and inactivation by moth antennae[J]. Nature, 1981, 293(5828): 161-163. |
| [10] | FIELD L M, PICKETT J A, WADHAMS L J. Molecular studies in insect olfaction[J]. Insect Mol. Biol., 2000, 9(6): 545-551. |
| [11] | VOGT R G. Molecular basis of pheromone detection in insects[M]//Comprehensive molecular insect science. Amsterdam: Elsevier, 2005: 753-803. |
| [12] | LIU S, GONG Z J, RAO X J, et al.. Identification of putative carboxylesterase and glutathione S-transferase genes from the antennae of the Chilo suppressalis (Lepidoptera: Pyralidae)[J/OL]. J. Insect Sci., 2015, 15(1): 103[2025-09-10]. . |
| [13] | LEITCH O, PAPANICOLAOU A, LENNARD C, et al.. Chemosensory genes identified in the antennal transcriptome of the blowfly Calliphora stygia [J/OL]. BMC Genomics, 2015, 16(1): 255[2025-09-10]. . |
| [14] | LEAL W S. Odorant reception in insects: roles of receptors, binding proteins, and degrading enzymes[J]. Annu. Rev. Entomol., 2013, 58: 373-391. |
| [15] | ISHIDA Y, LEAL W S. Chiral discrimination of the Japanese beetle sex pheromone and a behavioral antagonist by a pheromone-degrading enzyme[J]. Proc. Natl. Acad. Sci. USA, 2008, 105(26): 9076-9080. |
| [16] | RYBCZYNSKI R, VOGT R G, LERNER M R. Antennal-specific pheromone-degrading aldehyde oxidases from the moths Antheraea polyphemus and Bombyx mori [J]. J. Biol. Chem., 1990, 265(32): 19712-19715. |
| [17] | ROGERS M E, JANI M K, VOGT R G. An olfactory-specific glutathione-S-transferase in the Sphinx moth Manduca sexta [J]. J. Exp. Biol., 1999, 202(Pt 12): 1625-1637. |
| [18] | 田晓丽,许冬,刘毅,等.南亚果实蝇谷胱甘肽S-转移酶基因的鉴定及组织表达[J].西北农林科技大学学报(自然科学版),2025(10):122-131. |
| TIAN X L, XU D, LIU Y, et al.. Identification and tissue expression of glutathione S-transferase gene from fruit fly in South Asia[J]. J. Northwest AF Univ. Nat. Sci. Edit., 2025(10): 122-131. | |
| [19] | WOJTASEK H, LEAL W S. Conformational change in the pheromone-binding protein from Bombyx mori induced by pH and by interaction with membranes[J]. J. Biol. Chem., 1999, 274(43): 30950-30956. |
| [20] | MAÏBÈCHE-COISNE M, NIKONOV A A, ISHIDA Y, et al.. Pheromone anosmia in a scarab beetle induced by in vivo inhibition of a pheromone-degrading enzyme[J]. Proc. Natl. Acad. Sci. USA, 2004, 101(31): 11459-11464. |
| [21] | WEI H, TAN S, LI Z, et al.. Odorant degrading carboxylesterases modulate foraging and mating behaviors of Grapholita molesta [J/OL]. Chemosphere, 2021, 270: 128647[2025-09-10]. . |
| [22] | 杨斌,刘杨,王冰,等.害虫嗅觉行为调控技术的研究现状、机遇与挑战[J].中国科学基金,2020(4):441-446. |
| YANG B, LIU Y, WANG B, et al.. Research status, opportunities and challenges of olfactory behavior regulation technology of pests[J]. Fund. Res., 2020(4): 441-446. | |
| [23] | 柳峰,孙礼兵,姚金亮,等.不同性诱产品对小菜蛾诱集效果比较及对其种群动态监测[J].植物保护,2011,37(5):210-212. |
| LIU F, SUN L B, YAO J L, et al.. Evaluation of the trapping efficacy of different sex pheromone lures to Plutella xylostella(L.) and the monitoring of its population dynamics[J]. Plant Prot., 2011, 37(5): 210-212. | |
| [24] | YUAN W, RAO X, ZHONG B, et al.. Exploring the functional profiles of odorant binding proteins crucial for sensing key odorants in the new leaves of coconut palms in Rhynchophorus ferrugineus [J/OL]. Int. J. Biol. Macromol., 2024, 261(Pt 2): 129852[2025-09-10]. . |
| [25] | 赵莹.印度谷螟(Plodia interpunctella)产卵相关气味受体基因的鉴定与功能研究[D].长春:东北师范大学,2022. |
| [26] | XIA D, ZHENG R, HUANG J, et al.. Identification and functional analysis of glutathione S-transferases from Sitophilus zeamais in olfactory organ[J/OL]. Insects, 2022, 13(3): 259[2025-09-10]. . |
| [27] | OAKESHOTT J G, JOHNSON R M, BERENBAUM M R, et al.. Metabolic enzymes associated with xenobiotic and chemosensory responses in Nasonia vitripennis [J]. Insect Mol. Biol., 2010, 19(S1): 147-163. |
| [28] | TSUBOTA T, SHIOTSUKI T. Genomic analysis of carboxyl/cholinesterase genes in the silkworm Bombyx mori [J/OL]. BMC Genomics, 2010, 11: 377[2025-09-10]. . |
| [29] | 蒋秀云.大蜡螟嗅觉相关基因鉴定、序列特征与表达模式分析[D].合肥:安徽农业大学,2021. |
| [30] | 沈怿丹.亚洲玉米螟气味降解酶基因鉴定及谷胱甘肽转移酶OfurGSTd1功能分析[D].合肥:安徽农业大学,2020. |
| [31] | 张宇星.稻纵卷叶螟气味降解酶与茶尺蠖感觉神经元膜蛋白基因鉴定及表达谱分析[D].合肥:安徽农业大学,2019. |
| [32] | 王炜.苹果蠹蛾Delta和Epsilon家族GSTs基因的鉴定及功能研究[D].沈阳:沈阳农业大学,2019. |
| [33] | 马国强,高俊仙,廉梅霞.短角外斑腿蝗GST基因的鉴定及其杀虫剂敏感性[J].西北农林科技大学学报(自然科学版),2020,48(6):99-105. |
| MA G Q, GAO J X, LIAN M X. Identification of glutathione S-transferase(GST)gene family from Xenocatantops humilis Brachycerus and its insecticide sensitivity[J]. J. Northwest AF Univ. Nat. Sci. Ed., 2020, 48(6): 99-105. | |
| [34] | GAO H, DAI L, FU D, et al.. Isolation, expression profiling, and regulation via host allelochemicals of 16 glutathione S-transferases in the Chinese white pine beetle, Dendroctonus armandi [J/OL]. Front. Physiol., 2020, 11: 546592[2025-09-10]. . |
| [35] | HU F, YE K, TU X F, et al.. Identification and expression profiles of twenty-six glutathione S-transferase genes from rice weevil, Sitophilus oryzae (Coleoptera: Curculionidae)[J]. Int. J. Biol. Macromol., 2018, 120(Pt A): 1063-1071. |
| [36] | ZHU F, MOURAL T W, SHAH K, et al.. Integrated analysis of cytochrome P450 gene superfamily in the red flour beetle, Tribolium castaneum [J/OL]. BMC Genomics, 2013, 14: 174[2025-09-10]. . |
| [37] | 俞丽英.小菜蛾细胞色素P450基因的全基因组鉴定及其进化与表达模式[D].福州:福建农林大学,2014. |
| [38] | CHERTEMPS T, FRANÇOIS A, DURAND N, et al.. A carboxylesterase, Esterase-6, modulates sensory physiological and behavioral response dynamics to pheromone in Drosophila [J/OL]. BMC Biol., 2012, 10: 56[2025-09-10]. . |
| [39] | OAKESHOTT J G, CLAUDIANOS C, CAMPBELL P M, et al.. Biochemical genetics and genomics of insect esterases[M]// Reference Module in Life Sciences. Amsterdam: Elsevier, 2019 |
| [40] | ZHANG Y N, LI J B, HE P, et al.. Molecular identification and expression patterns of carboxylesterase genes based on transcriptome analysis of the common cutworm, Spodoptera litura (Lepidoptera: Noctuidae)[J]. J. Asia Pac. Entomol., 2016, 19(4): 989-994. |
| [41] | DURAND N, CAROT-SANS G, BOZZOLAN F, et al.. Degradation of pheromone and plant volatile components by a same odorant-degrading enzyme in the cotton leafworm, Spodoptera littoralis [J/OL]. PLoS ONE, 2011, 6(12): e29147[2025-09-10]. . |
| [42] | HE P, ZHANG Y N, LI Z Q, et al.. An antennae-enriched carboxylesterase from Spodoptera exigua displays degradation activity in both plant volatiles and female sex pheromones[J]. Insect Mol. Biol., 2014, 23(4): 475-486. |
| [43] | HE P, MANG D Z, WANG H, et al.. Molecular characterization and functional analysis of a novel candidate of cuticle carboxylesterase in Spodoptera exigua degradating sex pheromones and plant volatile esters[J]. Pestic. Biochem. Physiol., 2020, 163: 227-234. |
| [44] | LÖFSTEDT C, VAN DER PERS J N C, LÖFQVIST J, et al.. Sex pheromone of the cone pyralid Dioryctria abietella [J]. Entomol. Exp. Appl., 1983, 34(1): 20-26. |
| [45] | HALL D R, FARMAN D, DOMÍNGUEZ J C, et al.. Female sex pheromone of the cone moth, Dioryctria mendacella: investigation of synergism between type Ⅰ and type Ⅱ pheromone components[J]. J. Chem. Ecol., 2017, 43(5): 433-442. |
| [46] | LEE S C, LEE J W, LEE D H, et al.. Identification of sex pheromone components of Korean Dioryctria abietella (Lepidoptera: Pyralidae) population and synergism of pheromone and pine cone volatile blends[J]. J. Econ. Entomol., 2022, 115(1): 178-186. |
| [47] | JIANG X C, DONG W X, CHEN B, et al.. Electrophysiological and oviposition responses of Asian corn borer, Ostrinia furnacalis (Lepidoptera: Crambidae), to compounds rinsed from the surfaces of sugarcane and maize leaves[J]. Eur. J. Entomol., 2015, 112(2): 295-301. |
| [48] | CHOO Y M, PELLETIER J, ATUNGULU E, et al.. Identification and characterization of an antennae-specific aldehyde oxidase from the navel orangeworm[J/OL]. PLoS ONE, 2013, 8(6): e67794[2025-09-10]. . |
| [49] | WANG M M, HE M, WANG H, et al.. A candidate aldehyde oxidase in the antennae of the diamondback moth, Plutella xylostella (L.), is potentially involved in the degradation of pheromones, plant-derived volatiles and the detoxification of xenobiotics[J/OL]. Pestic. Biochem. Physiol., 2021, 171: 104726[2025-09-10]. . |
| [50] | DING Y, ORTELLI F, ROSSITER L C, et al.. The Anopheles gambiae glutathione transferase supergene family: annotation, phylogeny and expression profiles[J/OL]. BMC Genomics, 2003, 4(1): 35[2025-09-10]. . |
| [51] | KETTERMAN A J, SAISAWANG C, WONGSANTICHON J. Insect glutathione transferases[J]. Drug Metab. Rev., 2011, 43(2): 253-265. |
| [52] | LIU Y, TIAN X, GUI L, et al.. Molecular and functional characterization of an antenna-enriched glutathione S-transferase BminGSTd3 involved in undecanol degradation in the Citrus fruit fly, Bactrocera minax (Enderlein) (Diptera Tephritidae)[J/OL]. Int. J. Biol. Macromol., 2024, 256(Pt 2): 128514[2025-09-10]. . |
| [53] | SCOTT J G. Insect cytochrome P450s: thinking beyond detoxification[J]. Recent Adv. Insect Physiol. Toxicol. Mol. Biol., 2008, 117-124. |
| [54] | 杨欢.光肩星天牛气味降解酶基因鉴定、表达及功能探究[D]. 北京:北京林业大学, 2021. |
| [55] | FEYEREISEN R. Insect CYP genes and P450 enzymes[M]//Insect molecular biology and biochemistry. Amsterdam: Elsevier, 2012: 236-316. |
| [56] | INGELMAN-SUNDBERG M. The human genome project and novel aspects of cytochrome P450 research[J]. Toxicol. Appl. Pharmacol., 2005, 207(2): 52-56. |
| [57] | FEYEREISEN R. Arthropod CYPomes illustrate the tempo and mode in P450 evolution[J]. Biochim. Biophys. Acta BBA Proteins Proteom., 2011, 1814(1): 19-28. |
| [58] | 朱克森,黄钧鸿,冯启理,等.草地贪夜蛾与斜纹夜蛾解毒相关基因的比较分析[J]. 环境昆虫学报, 2020, 42(2): 318-328. |
| ZHU K S, HUANG J H, FENG Q L, et al.. Comparative analysis of detoxification related genes in Spodoptera frugiperda and Spodoptera litura [J]. J. Environ. Entomol., 2020, 42(2): 318-328. | |
| [59] | CUI S, WANG L, MA L, et al.. P450-mediated detoxification of botanicals in insects[J]. Phytoparasitica, 2016, 44(5): 585-599. |
| [60] | KEELING C I, HENDERSON H, LI M, et al.. CYP345E2, an antenna-specific cytochrome P450 from the mountain pine beetle, Dendroctonus ponderosae Hopkins, catalyses the oxidation of pine host monoterpene volatiles[J]. Insect Biochem. Mol. Biol., 2013, 43(12): 1142-1151. |
| [61] | FENG B, ZHENG K, LI C, et al.. A cytochrome P450 gene plays a role in the recognition of sex pheromones in the tobacco cutworm, Spodoptera litura [J]. Insect Mol. Biol., 2017, 26(4): 369-382. |
| [1] | 万令飞, 潘文婷, 雍雨婷, 李元帅, 赵悦, 阎新龙. 空间转录组学在肝病研究中的应用进展[J]. 生物技术进展, 2025, 15(4): 645-654. |
| [2] | 蔡怡霏, 马叶子, 夏美娟, 刘翠翠, 王洪涛, 周家喜. 小鼠胚胎主动脉-性腺-中肾区与成年骨髓巨核细胞的分子特征分析[J]. 生物技术进展, 2025, 15(4): 702-710. |
| [3] | 海萨·艾也力汗null, 焦飞, 刘鸿, 呼德拉提·阿那斯null, 沈玉帮. 联合GWAS和转录组分析挖掘与白斑狗鱼耐高温性状关联的SNP[J]. 生物技术进展, 2025, 15(3): 432-445. |
| [4] | 刘卓颖, 周晓今, 黄燕丽, 逄森. 玉米盐胁迫和MeJA处理下的转录组联合分析[J]. 生物技术进展, 2025, 15(2): 263-275. |
| [5] | 万令飞, 潘文婷, 雍雨婷, 李元帅, 赵悦, 阎新龙. 单细胞转录组测序技术在肝纤维化中的研究进展[J]. 生物技术进展, 2024, 14(5): 793-804. |
| [6] | 马叶子, 夏美娟, 刘翠翠, 王洪涛, 周家喜. 新型冠状病毒奥密克戎感染下血小板转录组与蛋白质组的对比分析[J]. 生物技术进展, 2024, 14(4): 649-656. |
| [7] | 张慧, 刘愽愽, 李凤梅, 崔健. 生长素信号途径参与南瓜白粉菌胁迫的响应研究[J]. 生物技术进展, 2024, 14(3): 433-441. |
| [8] | 乔红梅. 红树科陆生植物竹节树的转录组分析[J]. 生物技术进展, 2024, 14(2): 228-236. |
| [9] | 王彩虹, 邵煜涵, 蒋梦媛. IL-17抑制铜绿假单胞菌感染肺上皮细胞的机制研究[J]. 生物技术进展, 2024, 14(2): 304-311. |
| [10] | 刘勤勤, 黄柏铭, 马叶子, 刘翠翠, 王洪涛, 周家喜. 急性感染条件下巨核细胞与血小板转录组变化的对比分析[J]. 生物技术进展, 2023, 13(3): 465-472. |
| [11] | 李敏敏, 夏美娟, 赵晶晶, 陈肖源, 刘翠翠, 苏培, 王洪涛, 周家喜. 人类胚胎期和成年期巨核细胞的分子特征比较[J]. 生物技术进展, 2022, 12(4): 577-583. |
| [12] | 李东翱, 刘慧泉, 王秦虎. 小麦响应禾谷镰刀菌侵染的转录组学研究进展[J]. 生物技术进展, 2021, 11(5): 610-617. |
| [13] | 柯杨, 马瑜, 朱海云, 张育辉. RNA‑seq技术在水生生物生态毒理学中的应用进展[J]. 生物技术进展, 2021, 11(4): 535-541. |
| [14] | 方汉顺,,谢永盾,曾伟伟,郭会君,熊宏春,赵林姝,古佳玉,徐延浩,刘录祥. 小麦矮秆突变体jm22d响应赤霉素处理的转录组学分析[J]. 生物技术进展, 2020, 10(5): 503-516. |
| [15] | 常文婧,赵彦艳. 激光捕获显微切割在肿瘤多组学研究中的应用进展[J]. 生物技术进展, 2020, 10(2): 130-136. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||